7.5 Vegetative compatibility
For genetically different hyphae to interact, to take advantage of heterokaryosis or the sexual cycle, for example, the process of hyphal anastomosis must be closely regulated so that the physiological and genetic advantages of heterokaryosis can be realised without hazard. And there are hazards: hyphal anastomosis carries the risk of exposure to contamination with alien genetic information from defective or harmful cell organelles, viruses or plasmids. But nuclear and cytoplasmic control strategies have very different requirements. To maximise the advantage of sexual reproduction the controls must ensure that the nuclei are genetically as different as possible. In contrast, safe operation of the cell requires that cytoplasms that are to mingle must be as similar as possible. These features are under the control of genetic systems that:
- regulate the ability of hyphae to fuse, generally called vegetative compatibility. The phenotype of vegetative compatibility (also called vegetative, somatic, or heterokaryon incompatibility) is formation of a joint heterokaryotic mycelium with vegetative compatibility as a self/nonself recognition process occurring when hyphae of the same species fuse;
- and genes called mating type factors that regulate the ability of nuclei that have been brought together by hyphal fusion to undergo karyogamy and meiosis, and the phenotype of compatible interaction between mating type factors is occurrence of sexual reproduction (details in Chapter 8);
-
Vegetative compatibility is different from, but has a controlling influence over, mating type function in terms of both population structure and genetic diversity.
Vegetative compatibility is controlled by one to several nuclear genes that limit completion of hyphal anastomosis between colonies to those that belong to the same vegetative compatibility group (usually abbreviated to v-c group). Members of a v-c group possess the same vegetative compatibility alleles. Hyphal anastomosis is promiscuous in fungi, but compatibility of the cytoplasms determines whether cytoplasmic exchange will progress beyond the first few hyphal compartments involved in the initial interaction. Since the intracellular test for self/non-self-recognition (that is, vegetative compatibility) occurs after anastomosis, this is called post-fusion incompatibility. If the colonies involved are not compatible the cells immediately involved in anastomosis are killed (Fig. 7) by a programmed cell death response (Paoletti & Clavé, 2007; Paoletti, 2016). This strategy prevents transfer of nuclei and other organelles between incompatible strains, but if the incompatibility reaction is slow, a virus or cytoplasmic plasmid may be communicated to adjacent undamaged cells before the incompatibility reaction kills the hyphal compartments where anastomosis occurred (Bidard et al., 2013).
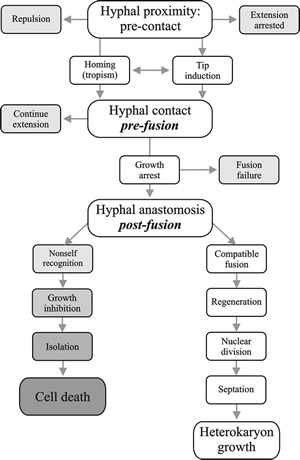 |
Fig. 7. Flow diagram illustrating the progress of hyphal interaction leading to operation of the vegetative compatibility systems. Recognition processes between hyphae take place at all three major steps: pre-contact hyphal proximity, pre-fusion hyphal contact and post-fusion self-non self recognition. Modified from Chapter 2 in Moore & Novak Frazer, 2002. |
The basis of a compatibility test carried out in the laboratory is that small pieces of the strains that are to be tested are placed side by side on the surface of an agar medium, and hyphal interactions in these ‘confrontations’ usually show phenotypes that imply self/nonself recognition. When the confrontations are incubated, leading hyphae may mingle, and hyphal anastomoses occur between their branches. If the confronting strains are compatible the heterokaryon may proliferate so that the whole mycelium becomes heterokaryotic; this is what happens in Neurospora crassa and Podospora anserina. Alternatively, in species such as Verticillium dahliae and Gibberella fujikuroi, nuclei do not migrate between cells and heterokaryosis is limited to the branches that grow out of the fusion cells.
If the colonies involved are not compatible, the fusion cells are killed (Fig. 7). Cell death resulting from vegetative incompatibility involves plugging of the septal pores, to compartmentalise dying hyphal segments; vacuolisation of the cytoplasm; DNA fragmentation; organelle degradation; and shrinkage of the plasma membrane from the cell wall. It is an internalised cell death, different from necrotic cell death, with many features in common with programmed cell death (PCD, or apoptosis) in other multicellular eukaryotes (Paoletti & Clavé, 2007; Paoletti, 2016).
| Table 3. Genes of filamentous fungi, involved in vegetative incompatibility, that have been cloned and characterised (Adapted from Moore & Novak Frazer, 2002). | |
| Neurospora crassa genes | |
| Mat A-1 | Mating type transcription regulator, contains region similar to Mat α1 of Saccharomyces cerevisiae. |
| Mat a-1 | Mating type transcription regulator with an HMG box (characteristic of High Mobility Group proteins, a class of proteins distinct from histones which are found in chromatin and represent a subclass of the non-histone proteins; the HMG proteins function in gene regulation and maintenance of chromosome structure). |
| het-c | Signal peptide (involved in endoplasmic reticulum targeting of secreted proteins) with glycine-rich repeats. |
| het-6 | Region of similarity to tol and het-e (of P. anserina). |
| un-24 | Large subunit of type I ribonucleotide reductase. |
| tol | Features a coiled-coil, leucine-rich repeat (a protein conformation found in extracellular matrix molecules), has regions similar to sequences in het-e (of P. anserina) and het-6. |
| Podospora anserina genes | |
| het-c | Glycolipid transfer protein (glycolipids are involved in cell to cell interactions). |
| het-e | GTP-binding domain, region with similarity to tol and het-6 of N. crassa. |
| het-s | Prion-like protein (abnormally-folded variant can infectively communicate its abnormal conformation to normal proteins which then form aggregates). |
| idi-2 | Signal peptide, induced by het-R/V incompatibility. |
| idi-1, idi-3 | Signal peptide, induced by nonallelic incompatibility. |
| mod-A | SH3-binding domain (src homology domain 3; a protein domain of about 50 amino acid residues present in proteins involved in signal transduction, and also in a number of cytoskeletal proteins, generally involved in protein-protein interactions). |
| mod-D | α-subunit of G-protein with GTP binding (such proteins are involved in signal transduction in eukaryotic cells), modifier of het-C/E incompatibility. |
| mod-E | Heat-shock protein (belongs to the Hsp90 family of 90 kDa polypeptides with ATPase activity which are essential for the viability of yeast cells and found in association with many regulatory proteins in eukaryotes, like steroid receptors and protein kinases), modifier of het-R/V incompatibility. |
| pspA | Vacuolar serine protease, induced by nonallelic incompatibility. |
Vegetative compatibility (also called vegetative or somatic incompatibility, is now increasingly being called heterokaryon incompatibility or HI) will prevent formation of a heterokaryon unless the strains belong to the same v-c group. You must cope with a variety of terms that were applied to this research through the 20th century; half of which was done by ‘glass half-empty people’ who knew they were researching incompatibility, and the other half by ‘glass half-full people’ who were researching compatibility.
Incompatibility between strains in a confrontation is caused by genetic differences between the two individuals at specific gene loci, which are called het (for heterokaryon) or vic (for vegetative incompatibility) loci, although once the major genes were identified several others that affected or otherwise modified their expression were also identified and given other descriptive names (Table 3).
There are usually about 10 het loci, but the number varies from one species to another: there are at least 11 het loci in Neurospora crassa, 9 in Podospora anserina, 8 in Aspergillus nidulans, and 7 in Cryphonectria parasitica. At the time of writing, only genes from the main models for the study of incompatibility, Neurospora (Hall et al., 2010) and Podospora (Bidard et al., 2013) have been cloned.
These genes encode STAND proteins (STAND proteins are Signal Transducing ATPases with Numerous Domains), their domains comprising:
- a carboxy-terminal WD-repeat domain (which features a sequence with a high frequency of tryptophan (symbol W) and aspartic acid (symbol D) pairs and named for the single-letter symbols commonly used in sequence data);
- a central NACHT nucleotide binding domain (the name NACHT is derived from the four animal and fungal proteins which initially defined the unique features of this domain [specifically: the Neuronal Apoptosis inhibitory protein, the major histocompatibility Complex transcription activator, the Podospora anserina incompatibility locus protein HET-E, and Telomerase-associated protein]; the NACHT domain has NTPase activity and preferentially binds GTP or ATP;
- and an N-terminal fungus specific HET domain, which is a cell death execution domain found in many proteins encoded by fungal incompatibility genes (Paoletti & Clavé, 2007; Paoletti, 2016).
Not surprisingly, given the range of controlling elements represented by these HI proteins, the incompatibility response involves massive changes to the transcriptome (the spectrum of messenger RNA molecules expressed from the genes of the organism). 2,231 genes were up-regulated by a factor 2 or more, and 2,441 genes were down-regulated during the incompatibility reaction in Podospora (Bidard et al., 2013). There was a significant overlap between regulated genes during incompatibility in Podospora anserina and Neurospora crassa, indicating similarities in the incompatibility responses in these two species. Some of the transcriptome changes observed during the incompatibility reaction mimic the impact of the host plant on plant pathogenic fungi, so it may not be surprising that the vegetative incompatibility groups into which strains of Rhizoctonia solani can be divided differ in their pathogenicities to the plant host.
Neurospora crassa is similar to Aspergillus in that heterokaryon formation requires genetic identity at all het genes. One of these genes is the mating type locus of N. crassa, and although this is unusual, association between mating-type and vegetative incompatibility is not restricted to N. crassa, but has been reported in Ascobolus stercorarius, A. heterothallicus, and Sordaria brevicollis. Usually, two different alleles of a het gene are found in wild type isolates, although het loci with more than two alleles have been found (for further details see chapter 2 in Moore & Novak Frazer, 2002).
In Neurospora crassa, heterokaryons made between strains of opposite mating type grow slowly and have an irregular colony outline as compared with the rapid, uniform growth of heterokaryons between strains with the same mating type. Evidently the mating type gene of Neurospora crassa controls both sexual compatibility and heterokaryon compatibility, although the former requires that the mating types are different, and the latter requires that the mating types are identical. It seems that nuclei of opposite mating type do not readily coexist in vegetative hyphae of Neurospora crassa. Aggressive maintenance of individuality between mates is neither unusual nor difficult to understand; in our own species, allegedly, men are from Mars, women from Venus. In Neurospora crassa, the molecular basis of this mating aggression is that the MATA-1 and MATa-1 mating polypeptides (detailed in Chapter 8, Section 8.3) encode transcription regulators that specify different cell types in the sexual phase, but they are lethal when expressed together in a vegetative cell. The mating function of MAT a-1 depends on its DNA-binding ability, but this is not needed for the vegetative incompatibility function. So, different functional domains of the polypeptide serve these two different activities of the mating type idiomorphs (Wang et al., 2012).
Heterokaryons made between N. crassa strains of the same mating type (and the same het genotype) have nuclear ratios close to 1: 1, full cytoplasmic continuity, and they also produce up to 30% heterokaryotic conidia. In the incompatible heterokaryon confrontations pores in the septa of any cells that do fuse become blocked, and the cytoplasm becomes granular, then vacuolated, and finally dies. When such cytoplasm, or even a phosphate-buffer extract of it, was injected into a different strain, the same degenerative changes resulted. The activity of the extract was associated with its proteins, demonstrating that heterokaryon compatibility self/non self recognition depends on the protein products of the het genes.
When two incompatible colonies of Podospora anserina meet, hyphal fusion is followed by death of the fused cells and consequent absence of pigment, so a clear zone forms between the colonies called a barrage. The barrage is due to vegetative incompatibility, but the colonies might still be sexually compatible, and if they are compatible, a line of perithecia can be formed on each side of the barrage because although fused vegetative cells are killed, lethality does not extend to fused sexual cells. The vegetative incompatibility genes are probably regulators of enzymes that trigger the cell senescence and death of the incompatible fusions, and the mating type factors probably protect sexually-compatible cells from the adverse effects of vegetative incompatibility genes.
Several het loci of P. anserina have been characterised, but the symbols have been assigned independently in the different fungi; that is, het-c in P. anserinahas no relationship to the het-c of N. crassa. Just as in N. crassa, the P. anserina het loci encode varied gene products (Table 2); the het-s gene product behaves like a prion protein. A prion is a ‘proteinaceous infectious particle’, a cellular protein that can assume an abnormal conformation that is infectious in the sense that it can convert the normal form of the protein into the abnormal (see Section 7.9, below). Hyphal anastomosis between het-s and the neutral het-s* strain results in the cytoplasmic transmission and infectious propagation of the het-s phenotype.
Although the het loci encode very different gene products, three regions of similarity can be detected between predicted products of the het-6 locus and the tol locus of Neurospora crassa, and the predicted product of the het-e locus of Podospora anserina. These regions might represent domains necessary for some aspect of vegetative incompatibility in which all three of these het loci are involved. Alleles of het-c that are found in N. crassa are present in other Neurospora species and related genera, indicating there was a common ancestor and conservation during evolution of this sequence. However, despite this indication that there may be some underlying similarity in function, a het locus from one species does not necessarily confer vegetative incompatibility in a different species.
Updated July, 2019
