5.11 The endomembrane systems
We have already mentioned the membrane-bounded organelles mitochondria and chloroplasts, which are the power generators for the cell, but eukaryotes depend on several other membrane systems that deserve reference.
Mitochondria contain their own genome separate from the nuclear genome, and their key role is to manage the flow of oxygen atoms, protons and electrons in the process of respiration. The purpose is to generate compounds (particularly ATP and NADH) containing chemical energy that can be used for useful work elsewhere.
Resources Box CLICK HERE to visit a YouTube animation describing the Mitochondrial Electron Transport Chain
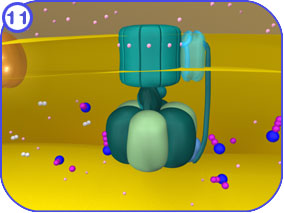
CLICK HERE to visit a YouTube animation describing ATP synthase powered by a hydrogen gradient
These animations belong to the Virtual Cell Animation Collection produced by the Molecular and Cellular Biology Learning Center at North Dakota State University http://vcell.ndsu.nodak.edu/animations/home.htm
|
Chloroplasts are the photosynthetic counterparts of mitochondria, using the photolysis of water to generate protons and electrons that are captured into chemical energy (mainly as ATP and NADPH). Organelles in plants that are modified chloroplasts are broadly called plastids, and are often involved in nutrient storage. Since both mitochondria and chloroplasts contain their own genomes, they are thought to have once been separate organisms, which formed symbiotic relationships at some stage in the evolution of eukaryotic cells. The most convincing version of this ‘endosymbiosis theory’ (Margulis, 2004) suggests a series of symbiotic relationships being established between prokaryotic partners: the mitochondria of eukaryotes evolving from aerobic bacteria living within a host cell; chloroplasts of eukaryotes evolving from endosymbiotic cyanobacteria; eukaryotic cilia and flagella arising from endosymbiotic spirochetes (Martin et al., 2015).
The other main components of the eukaryotic cell’s collection of single-layer endomembranes are the endoplasmic reticulum and the Golgi apparatus, both of which are responsible for managing the macromolecules of the cell. Endoplasmic reticulum (ER) is a transport network within the cell that sorts molecules intended for specific modifications and/or destinations. There are two forms of ER: rough ER has ribosomes docked on its cytoplasmic surface, anchored there because the polypeptides they are synthesising carried a signal sequence that directed them to the ER translocon [Section 5.10; CLICK HERE to see it now], and the smooth ER, which lacks docked ribosome. The ER is a system of membrane-bounded channels and sacs (called cisternae) connected to the double-layered nuclear envelope, which makes, processes, and transports chemical compounds for use throughout the cell, and provides a link between nucleus and cytoplasm.
The Golgi apparatus was one of the first intracellular organelles to be visualised, being observed by several workers at the end of the 19th century, though one called Camillo Golgi was the first to publish the observation. Golgi apparatus is another system of membrane-enclosed compartments, which modifies molecules made in the endoplasmic reticulum and prepares them for export to the outside of the cell.
This description implies that there is a constant traffic between the two membrane systems, and this is the case, transport or carrier vesicles bud from the endoplasmic reticulum membranes and selectively fuse with the Golgi apparatus (and there is a reverse, or retrograde, traffic too). These events require specific protein-protein interactions. Firstly, the vesicle forms through assembly of a protein coat on the cytoplasmic surface of the ER membrane which establishes curvature in the membrane and determines which membrane proteins are accepted by the vesicle. Secondly, and after transport, pairing of vesicle proteins with proteins in the Golgi membrane allows the vesicle and target membranes to fuse so that the cargo is deposited into the Golgi apparatus. Vesicle traffic from the ER to the Golgi involves vesicles coated with ‘coatomer protein’ COPII. A different vesicle coat protein complex (COPI) is responsible for transport from Golgi to ER, and also for transport between Golgi compartments (Day et al., 2013).
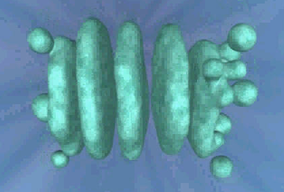
COPII proteins were first identified by genetic studies with yeast, but mammalian counterparts to the yeast genes were soon identified and shown to serve the same functions, so this is another highly conserved eukaryotic process. This molecular/genetic evidence is significant because there is a considerable difference in the morphology of the Golgi apparatus between fungi and mammals. The Golgi body in mammals (as originally observed in the cytoplasm of neurons by Golgi) is a stack of flattened large vesicles with swollen peripheral regions (the cisternae, often called dictyosomes by plant cell biologists), which are linked together by tubular connections and surrounded by numerous smaller spherical vesicles which have budded off from the cisternae. There are usually between five and eight cisternae in the stack but as many as sixty have been observed in some unicellular flagellates. Stacked Golgi bodies like this are not observed in plants or fungi. Most plant cells have hundreds of individual Golgi dictyosomes distributed throughout the cytoplasm. Fungi also have individual dictyosomes throughout the hypha but these are organised and consequently function so as to supply all the materials needed for rapid hyphal tip extension (Harris, 2013).
Such a complicated structure is obviously functionally differentiated and it’s easier to describe, and understand, the stacked Golgi apparatus of animals. Obviously, a stack of anything has a top and a bottom and it turns out that the stack of cisternae is functionally differentiated in that one end is the entry face, called the cis-Golgi, at which vesicles are received, the other end is the trans-Golgi, from which export vesicles leave. The medial (or middle) saccules are between the two, and both cis and trans are associated with networks of tubular and small cisternal structures, which are called the cis-Golgi network (CGN) and trans-Golgi network (TGN).
Resources Box CLICK HERE to visit a YouTube animation describing Protein Trafficking in the Golgi apparatus
These animations illustrate the animal Golgi comprised of stacked cisternae. This is not found in fungi, but fungi do carry out all these dynamic Golgi functions, though the dictyosomes are scattered through the cytoplasm and are not stacked into the historical 'Golgi body'. CLICK HERE to visit a YouTube animation describing Protein Modification within the Golgi apparatus
These animations belong to the Virtual Cell Animation Collection produced by the Molecular and Cellular Biology Learning Center at North Dakota State University http://vcell.ndsu.nodak.edu/animations/home.htm
|
The main functions of the Golgi apparatus are to modify, sort, package and transport substances formed by the ER. These reach the Golgi stack at its cis-side via vesicles budded-off the ER. Enzymes within the Golgi cisternae modify proteins by the addition of carbohydrates (glycosylation), sulfate (sulfation) and phosphate (phosphorylation). To achieve this the Golgi also imports substrates like phosphate and sulfate donors and nucleotide sugars from the cytoplasm. Proteins are also labelled with signal sequences to specify their final destination; for example, adding a mannose-6-phosphate labels proteins destined for lysosomes. The animal cell Golgi is also important in the synthesis of proteoglycans needed for the animal cell extracellular matrix. It is also a major site of carbohydrate synthesis.
The products of these modifications are packaged in membrane-bounded vesicles at the periphery of the trans-Golgi network that are targeted to the appropriate membrane somewhere else in the cell, such as lysosomes, peroxisomes and plasma membrane (for excretion from the cell). There is a flow in the other (retrograde) direction, too. Some of the product vesicles that emerge from the trans-Golgi network return ER-resident proteins to the ER and export Golgi-modified products for distribution by the ER.
In addition, we should appreciate that the cell plasma membrane (plasmalemma) is a membrane organelle in its own right and it is capable of both endocytosis (uptake) and exocytosis (secretion). The cell plasma membrane is made from a double layer of phospholipids. Because of its polar nature, the phosphatidyl ‘head’ of a phospholipid is hydrophilic (attracted to water) and the lipid tails are hydrophobic but lipophilic. Existing in an aqueous environment, the phospholipid bilayer is stable with the phosphatidyl heads on the two outer surfaces with the lipid tails together forming a lipophilic microenvironment between them. This forms a very fluid membrane that has a variety of protein molecules embedded within it acting as channels, transporters and receptors for regulation of the exchange and contact with the exterior of the cell.
Large molecules cannot pass through this membrane without active assistance. The process that imports these is called endocytosis. There are specialised regions called coated pits on the outside of the membrane. The ‘coat’ is a protein called clathrin, which can form a localised polyhedral lattice on the plasma membrane to cause an invagination of the membrane with the result that the imported cargo is drawn into the cytoplasm in a vesicle. This mechanism is used for uptake of essential metabolites, uptake of some regulators and growth factors, or uptake of previously-exported molecules for recycling. The resultant endocytic vesicles are transported to the Golgi apparatus and the ER where they fuse with Golgi and ER endosomes. After vesicle fusion the coat detaches and may be reused and the membrane that formed the invagination is ultimately returned to the cell surface. A similar clathrin-coating process also buds membrane segments from the trans-Golgi network.
The small transport vesicles we have already mentioned are defined by their coat proteins: COPII-coated vesicles allow export from the ER, COPI vesicles shuttle proteins between Golgi and ER, and clathrin-coated vesicles mediate transport from the trans-Golgi network and endocytic transport from the plasma membrane. These organised pathways require pairs of membranes to specifically recognise one another and subsequently fuse. Different targeting reactions involve distinct protein complexes that act to mark the target organelle for incoming vesicles (Faini et al., 2013). This is the responsibility of SNARE proteins (SNARE is an acronym derived from ‘soluble N-ethylmaleimide sensitive factor attachment protein receptor’).
These are are integral membrane proteins found predominantly on vesicles (v-SNAREs incorporated into the membranes of transport vesicles during budding) or target membranes (t-SNAREs located in the membranes of target compartments). SNAREs are fusion proteins; it is a large family of proteins with more than 60 members in yeast. Initial recognition between a vesicle and a target membrane is the responsibility of another group of proteins called tethering factors, which are large fibrous proteins that can span relatively long distances (>200 nm) between vesicle and target membrane, so they may form a molecular ‘fishing net’ to catch relevant vesicles. Tethering factors bind membranes together prior to the interaction of v-SNAREs and t-SNAREs across the membrane junction. The SNAREs form a complex that extends across both membranes and then ‘zippers’ the two membranes together (Wang et al., 2017).
The previous paragraphs describe the animal Golgi comprised of stacked cisternae. This is not found in fungi, but it is not true to say, as some accounts do, that ‘fungi lack Golgi dictyosomes’. Fungi do carry out all the Golgi functions, but the dictyosomes are scattered through the cytoplasm and are not stacked into the historical Golgi body (Day et al., 2013; Pantazopoulou, 2016). Cisternal (or dictyosome) stacking is an animal characteristic, but the stack consists of a collection of cisternae at different stages of maturation, with the youngest at the cis-side and most mature at the trans-side. As the trans-cisternae mature they 'dissolve' into vesicles and are replaced by the next-most-mature from the stack, and a new cisterna is assembled on the cis-side. This dynamic production line is well illustrated in the animation Protein Trafficking in the Golgi apparatus.
In plants and fungi the dictyosomes go through essentially the same maturation processes without being collected together into a stack.
Fungi have a main central vacuole as their principal degradative compartment and there are several pathways to the vacuole that supply it with hydrolytic enzymes, two from the trans-Golgi network, and a cytoplasm-to-vacuole pathway. There is a further range of vacuolar import pathways for substrate delivery: including endocytosis from the cell surface delivering metabolites in bulk as well as specific components, and regular transport of proteins destined for recycling in the vacuole. For the most part, flow to the vacuole happens in response to specific environmental stimuli; the main vacuole may accumulate any of a wide range of metabolic end-products varying from nutrient stores all the way to waste materials. Each of these accumulation, transport and delivery events depends on cellular machinery sensing the environmental signal(s), activating appropriate metabolic pathways and designating the appropriate transport mechanisms and targets (Veses et al., 2008; Richards et al., 2010, 2012; Tong et al., 2016).
The network of vesicle traffic in the growing hypha was first visualised using fluorescent dyes that insert into the outer layer of the plasma membrane (Fischer-Parton et al., 2000). Within 30 seconds of adding the dye to hyphae a cloud of fluorescent endocytic vesicles appears within the cytoplasm. Although components of the endocytic pathway have been characterised by molecular and genetic means, uptake of the dye is a clear demonstration that the full endocytosis process does occur in filamentous fungi.
The summarised model that results from the observations (Fig. 5) shows that the next stained organelles that could be visualised were small and roughly spherical endosomes. Subsequently, in subapical compartments, the next obviously stained organelle was the large main vacuole. This sequence is similar to observations of budding yeast, where staining of vacuole membranes followed that of endosomes. In hyphae the vacuolar system consists of an extensive tubular network in addition to the large roughly spherical vacuoles. Both the Golgi and ER of fungal hyphae would become stained via pathways connecting the endosomal system, Golgi and ER, which are known to occur in budding yeast.
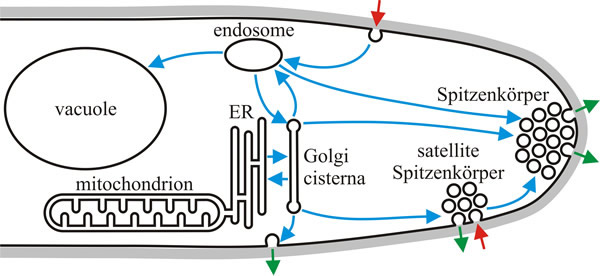 |
| Fig. 5. A simplified hypothetical model of the organisation of the vesicle trafficking network in a growing hypha based on the pattern of fluorescent dye staining. Intracellular trafficking is shown with blue arrows, exocytosis with green arrows and endocytosis with red arrows. Plasma membrane and other membrane compartments are shown in black with the hyphal cell wall shown in grey. ER = endoplasmic reticulum. Redrawn and modified from Fischer-Parton et al., 2000. |
The model shown in Fig. 5 also introduces a membranous organelle that is a characteristic feature in fungi with true hyphae: an apical cluster of vesicles and cytoskeletal elements, which plays a crucial role in hyphal tip extension growth and is known by its German name, the Spitzenkörper (= ‘apical body’). The Spitzenkörper is the organising centre for hyphal extension and morphogenesis; present in actively growing tips but lost when extension ceases. It can be detected with phase contrast light microscopy in growing hyphae of Ascomycota and Basidiomycota. Some zygomycetes lack a recognisable Spitzenkörper, but do have a loose distribution of vesicles in the hyphal apex that may serve the same function. The structure of the Spitzenkörper, as seen by light microscopy, differs in detail between species, but is also dynamic and variable within a species. Even though it lacks a membrane boundary it is clearly a complex organelle, which is always adjacent to the site of polarised hyphal extension. It is comprised of vesicles of various kinds and sizes, microfilaments, microtubules, and ribosomes.
Extension of the hyphal tip requires polarised incorporation of plasma membrane and cell wall constituents into the growing apex. Apical extension of the hyphal tip is considered to depend on the supply of wall-building secretory vesicles generated by Golgi cisternae and discharged from a ‘vesicle supply centre’ which is the cytologically-visible Spitzenkörper. As Fig. 5 suggests, satellite Spitzenkörper(s) arise immediately beneath the plasma membrane a few micrometres behind the apex, before migrating towards and merging with the main Spitzenkörper to supply additional wall-building vesicles to the growing hyphal tip. In experiments with fluorescent dye the Spitzenkörper stained shortly after the endosomes suggesting that, as well as the Golgi, the endosomal system may supply materials to the Spitzenkörper. It is plausible that recycled plasma membrane, captured by endocytosis and sorted by the endosome(s), is among the material supplied to the Spitzenkörper by the vesicle flow (Harris et al., 2005; Virag & Harris, 2006; Steinberg, 2007b; Riquelme, 2013; Steinberg et al., 2017). Fluorescent probes also label vacuoles that form a tubular reticulum in hyphal tip cells, and short tubules that undergo sequences of characteristic movements and transformations to produce ring-like membrane structures not yet fully identified. The cell biology of the apical dome of the hypha is complex and dynamic (Zhuang et al., 2009), and the vacuolar system of the hypha exhibits multiple stress responses and environmental adaptations (Tong et al., 2016).
Updated July, 2019