11. Plausible molecular mechanisms
Epigenetics is not often discussed in relation to fungal development, and when it is the more restricted interpretation of epigenetic modification of the genome is the usual topic, with the implication that gene expression takes pride of place in morphogenesis. This comment is not intended to underestimate any role played by such mechanisms. There are a number of processes that silence genes in filamentous fungi (Irelan & Selker, 1996). One of these is transcriptional silencing of genes by methylation of cytosines in the DNA which seems to be common in fungi and plants (Meyer, 1996). Pre-meiotic methylation occurs during the sexual cycle in Ascobolus immersus and Coprinopsis cinerea to silence repeated sequences reversibly. The mechanism involves homologous pairing to sense the presence of duplications. Differences in methylation patterns have been detected between yeast-phase and filamentous growth in some dimorphic fungi (Reyna Lopez, Simpson & Ruiz-Herrera, 1997). It is this reversible differential DNA methylation which might be related to fungal morphogenesis. This is an attractive proposition in general theoretical terms (Bestor, Chandler & Feinberg, 1994). There is, however, also evidence that methylation plays a role in the control of chromosome behaviour during meiosis in Neurospora crassa (Foss et al., 1993). This specific possibility argues against DNA methylation being involved generally in morphogenesis. A similar conclusion is indicated by the facts that methylation occurs to a lesser extent in the mushroom Coprinopsis cinerea than it does in Ascobolus or Neurospora (Freedman & Pukkila, 1993), and that there was no evidence for development-related differences in methylation in Agaricus bisporus (Wilke & Wach, 1993).
Neither methylation nor any other single mechanism provides an adequate foundation for a developmental network at the gene expression level. My preference is for a more holistic approach to identifying factors with the potential to influence morphogenesis.
As stated earlier [view now], key words at each stage of development in fungi seem to be competence, induction and change. Competence is repeatedly encountered. Hyphae must be able to initiate the next step, but the next step is not inevitable. Competence may be genetic (e.g. mating types) but is primarily a physiological state. Induction is the process by which the competent tissue is exposed to conditions which overcome some block to progress and allow the next stage to proceed. Change occurs when the competent tissue is induced. The next stage always involves change in hyphal behaviour and physiology. This is usually quite drastic and represents an additional property to those already expressed. That is, each developmental step takes the tissue to a higher order of differentiation. In relation to earlier discussion in this chapter, the state of competence represents the establishment of a network in the sense of Trewavas (1986, and see below), but expanded to include the full range of physical and physiological components.
Differentiated hyphal cells require reinforcement of their differentiation 'instructions'. This reinforcement is part of the context within which they normally develop (that is, it is part of their network), but when removed from their normal environment most differentiated hyphae revert to the mode of differentiation which characterises vegetative hyphae. Hyphal differentiation is consequently an unbalanced process in comparison with vegetative hyphal growth. In most hyphal differentiation pathways the balance must be tipped in the direction of 'differentiation' by the local microenvironment which is, presumably, mainly defined by the local population of hyphae.
In overall summary of this section (Fig. 41), having multicellular morphogenesis compartmentalised into a collection of distinct developmental processes in fungi, the 'subroutines' discussed above which give rise to the ‘normal’ morphology when played out in their correct arrangement, permits the flexibility in expression that allows the fruit body to react to adverse conditions and still produce a crop of spores; this being the foundation of the tolerance of imprecision referred to above. The ultimate flexibility, of course, is that the differentiation process can be abandoned in favour of vegetative hyphal growth and reversion to the invasive mycelium ensues. A lesser level of flexibility may be that a particular function is carried out by an incompletely adapted cell type. Differentiation is influenced by the local environment and hyphae influence their own local environment by producing and exporting components of their extracellular matrix chemically tailored to produce the required ionic environment.
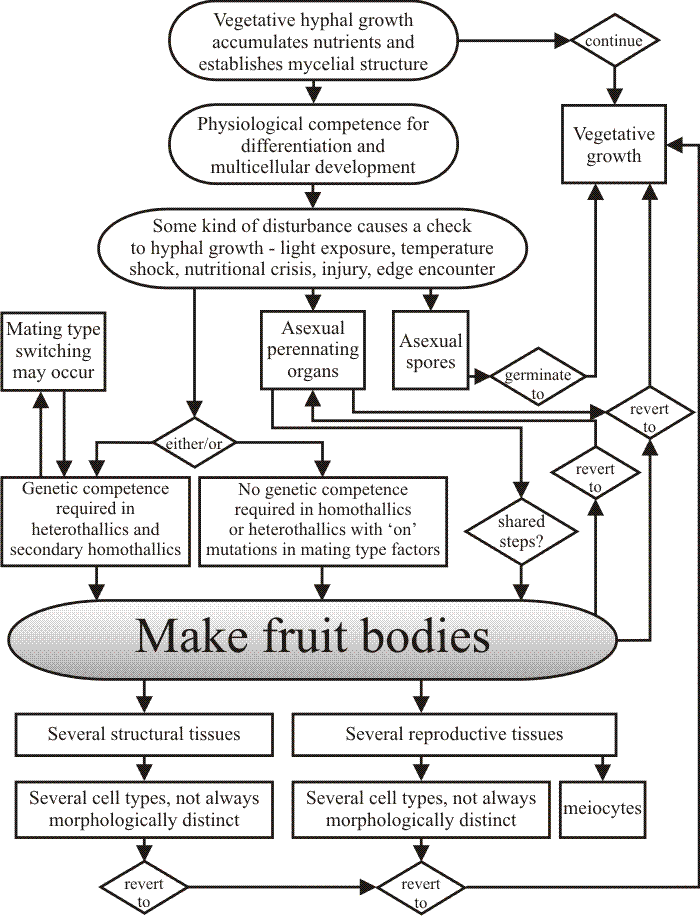 |
| Fig. 41. Flow chart showing a simplified view of the processes involved in development of fruit bodies and other multicellular structures in fungi. From Moore, 1998a. |
When it comes to searching for mechanisms which might control fungal morphogenesis there is no shortage of candidates. Homologues and analogues of all of the mechanisms known in animals and plants can be found in fungi. For control at the genetic level the mating type factors provide prime examples of transcriptional control elements able to regulate specific morphogenetic subroutines. The flow chart in Fig. 41 summarises all of these notions.
Principles of mushroom development. In many fungi hyphae differentiate from the vegetative form that ordinarily composes a mycelium and aggregate to form tissues of multihyphal structures. These may be linear organs (that emphasise parallel arrangements of hyphae), such as strands, rhizomorphs and fruit body stems; or globose masses (that emphasise interweaving of hyphae), such as sclerotia, fruit bodies and other sporulating structures of the larger Ascomycota and Basidiomycota. Fungal morphogenesis depends on a series of principles (Table 4), most of which differ from those that characterise the developmental processes of both animals and plants (Moore, 2005).
It is important to keep in mind the structures and patterns to which these deduced developmental principles contribute. The fundamental structure of the mushroom fruit body is summarised in Figs 42 and 43.
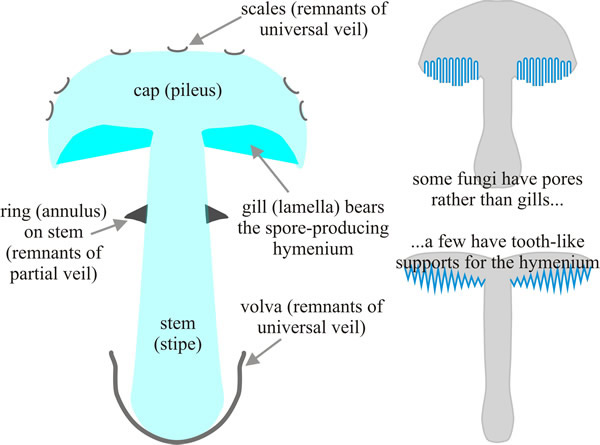 |
| Fig. 42. Main tissues of a mushroom. The mushroom is a spore-dispersal organ that is basically umbrella-shaped (and has the same function, to protect against rain, because proper spore release is prevented by liquid water), with the spore-bearing hymenium lining the under surface of a cap (also called pileus) and supported above the substratum by the stem (= stipe), shown in this diagrammatic vertical section. The mature mushroom may also bear remnants of thin tissues (veils) that protect the juvenile stages. Some fungi have a tubular or a toothed hymenium rather than one borne on gills; nevertheless, in all cases when the basidiospores are released they must fall vertically to escape from the cap for dispersal in the air currents below. Consequently these mushroom fruit bodies are exquisitely sensitive to the gravity vector and exhibit strong and rapid gravitropism. Illustration from chapter 12 in Moore, Robson & Trinci (2011). |
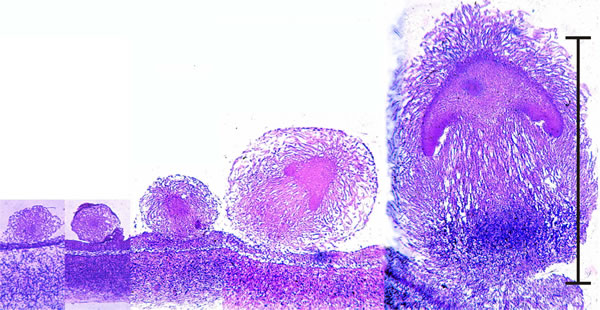 |
| Fig. 43. Early development of something like a mushroom looks very much like an embryonic process (scale bar = 1 mm), but remember that fungi are clonal organisms and these objects are fruit bodies, and many of them may be produced by an individual mycelium over an extended period of time. These images are photomicrographs of light microscope sections of successive stages in the very early development of the fruit body of the Ink Cap mushroom, Coprinopsis cinerea. The mature fruit body is approximately 100 mm tall, so this sequence covers just the first 1% of its developmental programme. The sections have been stained with the Periodic Acid-Schiff reagent, which stains polysaccharide accumulations blue-purple; in this case the accumulated polysaccharide, identified by other analyses, is glycogen. The object at extreme left is a large hyphal tuft, at second left is an initial (it shows some internal compaction and differentiation and can become either a sclerotium or a fruit body, depending on environmental conditions). Note that the third section is obviously differentiated into cap-like and stem-like structures, even though it is only 300 μm tall, and this is even more evident in the fourth section (700 μm tall), which has young gills but no gill cavity. At extreme right is a 1.2 mm tall fruit body primordium, in which the basic ‘body plan’ of the mushroom (see Fig. 42) is complete with clear demarcation into veil, pileipellis (= cap epidermis), cap, gills (with the beginnings of an annular gill cavity), and stem (with a distinct stem basal bulb which features heavy accumulations of glycogen). Illustration from chapter 12 in Moore, Robson & Trinci (2011). |
Most of these principles emerged from work which has already been described. However, Principle 5 (because hyphae grow only at their apex, global change to tropic reactions of all the hyphal tips in a structure is sufficient to generate basic fruit body shapes) was established from detailed mathematical models of morphogenesis and hyphal growth in Coprinopsis, which derived from work on gravitropism in the late 1990s that continued into the 21st century [view now].
| Table 4. The principles of fungal developmental biology | |
| Principle 1 | The fundamental cell biology of fungi on which development depends is that hyphae extend only at their apex, and cross walls form only at right angles to the long axis of the hypha. |
| Principle 2 | Fungal morphogenesis depends on the placement of hyphal branches. Increasing the number of growing tips by hyphal branching is the equivalent of cell proliferation in animals and plants. To proliferate, the hypha must branch, and to form an organised tissue the position of branch emergence and its direction of growth must be controlled. |
| Principle 3 | The molecular biology of the management of cell-to-cell interactions in fungi is completely different from that found in animals and plants. |
| Principle 4 | Fungal morphogenetic programmes are organised into developmental subroutines, which are integrated collections of genetic information that contribute to individual isolated features of the programme. Execution of all the developmental subroutines at the right time and in the right place results in a normal structure. |
| Principle 5 | Because hyphae grow only at their apex, global change to tropic reactions of all the hyphal tips in a structure is sufficient to generate basic fruit body shapes. |
| Principle 6 | Over localised spatial scales co-ordination is achieved by an inducer hypha regulating the behaviour of a surrounding knot of hyphae and/or branches (these are called Reijnders’ hyphal knots). |
| Principle 7 | The response of tissues to tropic signals and the response of Reijnders’ hyphal knots to their inducer hyphae, coupled with the absence of lateral contacts between fungal hyphae analogous to the plasmodesmata, gap junctions and cell processes that interconnect neighbouring cells in plant and animal tissues suggest that development in fungi is regulated by morphogens communicated mainly through the extracellular environment. |
| Principle 8 | Fungi can show extremes of cell differentiation in adjacent hyphal compartments even when pores in the cross wall appear to be open (as judged by transmission electron microscopy). |
| Principle 9 | Meiocytes appear to be the only hyphal cells that become committed to their developmental fate. Other highly differentiated cells retain totipotency - the ability to generate vegetative hyphal tips that grow out of the differentiated cell to re-establish a vegetative mycelium. |
| Principle 10 | In arriving at a morphogenetic structure and/or a state of differentiation, fungi are tolerant of considerable imprecision (= expression of fuzzy logic), which results in even the most abnormal fruit bodies (caused by errors in execution of the developmental subroutines) being still able to distribute viable spores, and poorly (or wrongly-) differentiated cells still serving a useful function. |
| Principle 11 | Mechanical interactions influence the form and shape of the whole fruit body as it inflates and matures, and often generate the shape with which we are most familiar. |
| From Moore, 2005. | |
Also, Principle 3 (the molecular biology of the management of cell-to-cell interactions in fungi is completely different from that found in animals and plants) emerged from an increasingly detailed series of genomic data-mining surveys that started in 2005 (Moore, Walsh & Robson, 2005) and continued through to 2009 (Moore & Meškauskas, 2006, Moore & Meškauskas, 2009).
Moore, Walsh & Robson (2005) initiated the study by searching a representative collection of filamentous fungal genomes (genomes of Coprinopsis cinerea,
Ustilago maydis, Cryptococcus neoformans, Phanerochaete
chrysosporium, Aspergillus nidulans, Neurospora crassa and
Aspergillus fumigatus) for gene sequences generally considered to be essential and highly conserved components of normal development in animals (specifically, sequences known as Notch, TGF-β, Wnt and Hedgehog). Genomes of the animals Caenorhabditis elegans (nematode) and Drosophila melanogaster (dipteran fruit fly) were searched as positive controls; and as an additional comparison, the genome of
the plant Arabidopsis thaliana (mouse-ear cress) was searched as a negative control for each sequence. The searches failed to reveal any homologies in the fungal genomes, and
Wnt, Hedgehog, Notch and TGF-β were all absent from Arabidopsis (though this plant has its own highly developed signalling pathways). We concluded that fungal and animal lineages diverged from their common opisthokont line well before the emergence of any multicellular arrangement, and that the unique cell biology of filamentous fungi has caused control of multicellular development in fungi to evolve in a radically different fashion from that in animals, and, probably, plants as well. A more comprehensive comparative analysis was done by Moore & Meškauskas in 2006, although the full search results were not published until three years later (Moore & Meškauskas, 2009). This was a fully comprehensive data-mining exercise, involving an estimated total of 590,000 similarity searches. It made use of software 'robots' or 'agents' released onto the Internet to search for homologies to polypeptide sequences assigned to the category ‘development’ in the Gene Ontology Consortium AmiGO database (www.geneontology.org) across all genomes available for all three of the crown group of eukaryote kingdoms. At the time this analysis was done there were 552 such developmental sequences and 1069 eukaryote genomes were available. It emerged that only 78 of the sequences were shared between all three Kingdoms, 72 were shared only between fungi and animals, 58 sequences were shared between plants and fungi, and 4 sequences were common only to the cellular slime mould Dictyostelium and fungal genomes.
No sequences were strictly fungus specific, but 68 occurred only in plants (Viridiplantae) and 239 occurred only in animals (Metazoa). Although some homology was indicated for a total of 219 fungal sequences, 143 (65%) of the matches returned were assigned probability values of only 5% and were categorised as weak similarities at best and not considered significant. The majority of the highly similar matches found in this survey proved to be between sequences involved in basic cell metabolism or essential eukaryotic cell processes (enzymes in common metabolic pathways, transcription regulators, binding proteins, receptors and membrane proteins). What were lacking were cross-kingdom similarities in the management processes that regulate multicellular development. These findings show categorically that there are no resemblances between the crown group of eukaryotic Kingdoms in the ways they control and regulate their developmental processes. Current understanding of phylogenetic relationships is that the major kingdoms of eukaryotes separated from one another at a stage prior to the multicellular grade of organisation. Consequently, in the course of their evolutionary history these very different organisms may have found different strategies to solve the same sorts of morphogenetic control problems. Unfortunately, this means, in particular, that we are currently totally ignorant of the way fungi regulate their multicellular development although there are some hints in the literature both old and new, at least about some plausible downstream control events.
Muraguchi, Takemaru & Kamada (1999) isolated over 1,000 developmental variants in fruiting using a homokaryotic fruiting strain (AmutBmut) of Coprinopsis cinerea and classified them into 8 groups (some of which are shown in Fig. 44): (1) knotless variants, which fail to form hyphal knots, the first visible sign of fruiting; (2) primordiumless variants, which form hyphal knots but fail to develop fruit body primordia; (3) maturationless variants, which form fruit body primordia but do not form mature fruit bodies; (4) elongationless variants, which form mature fruit bodies with short stems; (5) expansionless variants, which form mature fruit bodies with unexpanded caps; (6) sporeless variants, which fail to produce black basidiospores, resulting in fruit bodies with white caps after maturation; (7) compound type, which includes variants exhibiting several of the phenotypes described above; (8) others, including variants that produce a ‘dark stipe’ etiolated fruit body even under alternating light/dark growth conditions (wild type etiolated fruit bodies form only in continuous darkness).
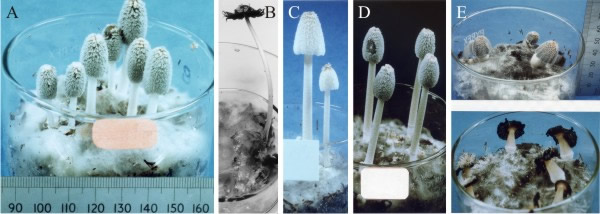 |
| Fig. 44. Developmental mutants of Coprinopsis cinerea. Shown from left to right are A and B the wild type (A mid-development, B fully mature and autolysing), and then C sporeless, D cap expansionless, and E stipe elongationless (immature at top, fully mature and autolysing below). The three mutants are all dominant and segregate in crosses as single genes. All cultures are contained in 9 cm diameter crystallising dishes. Photographs by Dr Junxia Ji. Illustration from chapter 12 in Moore, Robson & Trinci (2011). |
With the advent of reliable methods to isolate and sequence specific DNA fragments, several such mutants of Coprinopsis cinerea blocked at different stages in fruit body development have been identified and characterised. The gene functions include:
- cyclopropane fatty acid synthase (cfs1): the normal fruit body initiation pathway includes the dark-dependent formation of primary hyphal knots and their light-induced transition to the more compact secondary hyphal knots. The cfs1 mutant forms primary hyphal knots, but these fail to progress. The cfs1 gene encodes a protein similar to cyclopropane fatty acid synthase, a class of enzymes shown in prokaryotes and plants to convert membrane-bound unsaturated fatty acids into cyclopropane fatty acids. The Escherichia coli enzyme is thought to be involved in the long-term survival of non-growing cells and its expression can be associated with environmental stresses. In C. cinerea the enzyme may modify the hyphal membrane to adapt it to some function essential to the fruit body developmental pathway (Liu, Srivilai, Loos, Aebi & Kües, 2006).
- pseudoclamp connection formation (pcc1) gene has been shown to function in initial sexual development and to prevent clamp cell formation in AmutBmut homokaryons. Cloning and sequencing of pcc1 showed that it encodes an SRY-type HMG protein. SRY is the ‘sex-determining region Y’ sequence on the Y chromosome in placental mammals and marsupials, and HMG proteins interact directly with chromatin and recruit transcription factors to bind to enhancers. The sequence of the HMG box of pcc1 in Coprinopsis cinerea was very similar to mating type sequences of Podospora anserina, Schizosaccharomyces pombe, and Ustilago maydis. It is thought that pcc1 might play an important role in coordinating and enhancing the activities of the A and B mating type genes in C. cinerea (Murata, Fujii, Zolan, & Kamada, 1998).
- the clp1 (clampless) mutation of the AmutBmut strain blocks clamp connection formation. Formation of clamp connections is an essential step in mating type A-regulated sexual development. Sequencing the clp1 genomic DNA identified a sequence, interrupted by one intron, encoding a novel protein of 365 amino acids. The clp1 transcript was normally detected only in the presence of the HD1/HD2 heterodimer protein formed when homokaryons with compatible A genes are mated. Forced expression of clp1 by promoter replacements induced clamp development without the need for a compatible A gene combination. Expression of clp1 is necessary and sufficient for induction of the A-regulated pathway that leads to clamp connection development (Inada, Morimoto, Arima, Murata & Kamada, 2001).
- fungal galectins, cgl1 and cgl2 (β-galactose-binding proteins), are differentially regulated during fruit body formation. Expression of cgl2 was initiated during hyphal knot formation and was maintained to maturation; cgl1 was specifically expressed in primordia and mature fruit bodies. This is discussed in the section entitled Gravitropism as a morphogenetic model [VIEW NOW].
- dark stipe (dst) genes: (discussed in section 2 Stem and cap morphogenesis in Coprinopsis fruit bodies [VIEW NOW]). The mutations known as dst1 and dst2 have been cloned and sequenced and are thought to contribute to the blue light photoreceptor of Coprinopsis cinerea; dst2 may have a role in blue light perception with dst1 being responsible for communication of the photoreception signal.
- the ich1 (ichijiku) developmental mutant fails to develop a differentiated cap at the apex of the primordial stem in early stage fruit body primordia. The ich1 gene was mapped to chromosome XII and found to encode a novel protein of 1,353 amino acids. The amino-acid sequence contains nuclear targeting signals, which mark the protein for import into the nucleus, suggesting that the ich1 protein functions in the nucleus. Northern blots showed that the ich1 gene is specifically expressed in the cap of the wild type fruit body. No ich1 mRNA was detected in the ich1 mutant, indicating loss of the promoter region of ich1 in the mutant genome. Taken together, these data demonstrate that the ich1 gene product is essential for cap formation (Muraguchi & Kamada, 1998).
- Stem elongation mutations (eln): a dominant mutation of the elongationless2 (eln2) gene affects pattern formation in the fruit body primordia, which have disturbed tissue organisation and cell morphogenesis in the primordial stem and mature into fruit bodies with short stems. The eln2 locus was assigned to chromosome XII and shown to encode a novel type of microsomal cytochrome P450 enzyme (CYP502) which is constitutively expressed and presumably essential for the rapid energy metabolism demanded by stem elongation (Muraguchi & Kamada, 2000). The elongationless3 (eln3) mutant produces a fruit body in which the stem hardly elongates. In the wild type, elongation growth of cylindrical stem cells is responsible for stem elongation, and these are fully inflated to make side-to-side contact with one another (see discussion of stem mechanics in the section above entitled Quantitative analysis of cell size and distribution [VIEW NOW]), whereas in the mutant, the organisation of the stem tissue is disturbed with much space between stem cells. This disorganization of the stem tissue, together with reduced elongation of the stem cells, causes the mutant stem short and bulging. The eln3 gene sequence is predicted to encode a protein of 927 amino acids with a general glycosyltransferase domain and to be located in the plasma membrane. Transcription of the eln3 gene is specifically activated in rapidly elongating stems. The putative Eln3 enzyme is presumably necessary for the wall synthesis demanded by the inflation and elongation of stem cells discussed elsewhere [VIEW NOW] (Arima, Yamamoto, Hirata, Kawano & Kamada, 2004).
Although the genes listed immediately above are essential for their particular developmental processes, in general they cannot be described as major regulators of development. The most obvious candidates for the description ‘major regulators’ are the mating type factors of Coprinopsis cinerea. In C. cinerea, sexual development is under the control of the A and B mating type factors in the sense that in the wild type both A and B mating type factors must be compatible (that is, genetically different) for a dikaryon to form between two mating partners (Fig. 45) (see Chapters 7 and 8 in Moore, Robson & Trinci (2011) for a complete explanation).
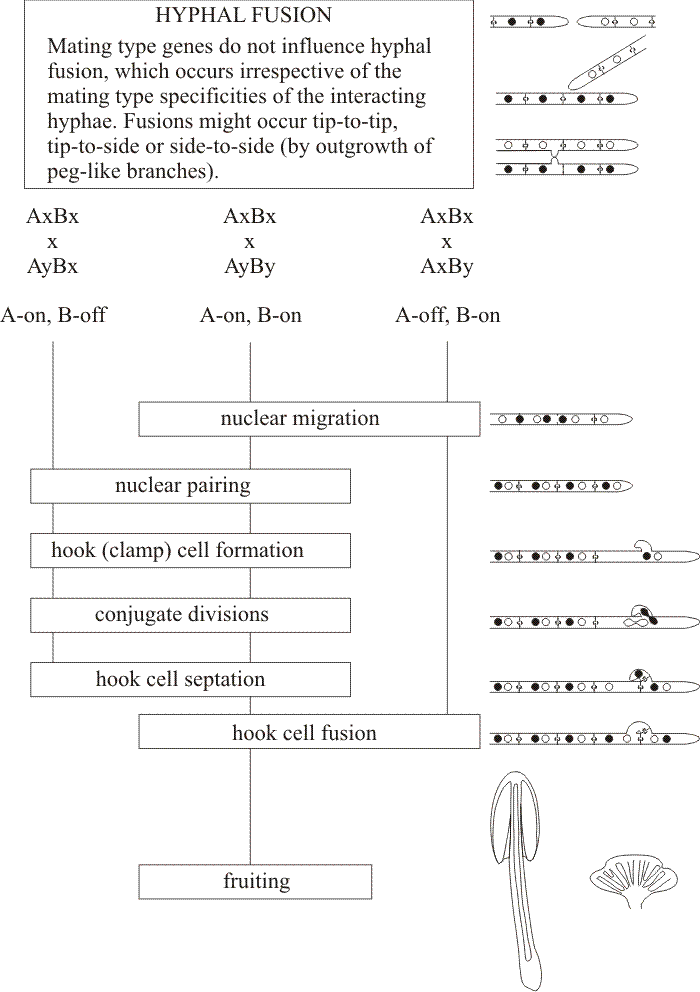 |
| Fig. 45. Flow chart diagram of A and B mating type factor activity in the Basidiomycota Coprinopsis cinerea and Schizophyllum commune. Many basidiomycetes have two unlinked mating type factors (designated A and B), an arrangement described as a bifactorial (because there are two factors) or tetrapolar (because the unlinked genes produce four progeny genotypes in every mating) incompatibility system. Compatibility requires that the two interacting mycelia carry different alleles, and both mating type factors must differ. The natural population contains many different mating types which behave in crosses as though they are multiple alleles at the two mating type loci. However, molecular analyses reveal that mating type chromosomal loci are composed of many mating type genes/gene pairs which is why the functional mating type factors are called ‘factors’ rather than ‘genes’. Compatible mating with formation of clamp connections and conjugate nuclei in the mated hyphae requires heterozygosity at both A and B (= A-on, B-on). Mating type factor A controls nuclear pairing, clamp connection formation and synchronised (conjugate) mitosis whereas mating type locus B controls nuclear migration and clamp connection tip fusion. Nuclei migrate through the existing mycelium after breakdown of septa between adjacent cells. Heterokaryons may be formed in matings which are homozygous for one of the mating type factors. When A factors are the same (A-off, B-on; = common-A heterokaryon), nuclear migration occurs but no clamp connections form. Mating of strains with the same B factor (A-on, B-off; = common-B heterokaryon) forms a heterokaryon only where the mated monokaryons meet because nuclear migration is blocked. Terminal cells of common-B heterokaryotic hyphae initiate clamp connections and nuclei divide but the hooked clamp connection cell fails to fuse with the subterminal cell of the hypha and its nucleus remains trapped. Diagram modified from Chiu & Moore (1998). |
Both of the mating type factors are complex genetic loci. In fact, each A locus of C. cinerea contains a variable number of genes, which are arranged in pairs encoding two dissimilar homeodomain proteins called HD1 and HD2 (the homeodomain is a protein structural domain that binds DNA or RNA and is thus commonly found in transcription factors). Sexual development is triggered by a dimerisation between HD1 and HD2 proteins from the different A mating type factors of the two nuclei of a compatible (dikaryotic) pairing. The A genes regulate conjugate division of the two nuclei from each mating partner and promote the formation of clamp connections (hyphal branch configurations at each septum required to maintain the dikaryotic state).
The B mating type factor contains three groups of genes which each code for a pheromone receptor and two pheromones. The B pheromone genes are all predicted to encode for lipopeptides, while the B pheromone receptors are typical G protein coupled receptors. The B genes regulate nuclear migration required for dikaryosis as well as the hyphal fusions that complete clamp connections. The model of pheromone function that has been developed for these filamentous basidiomycetes is that after anastomosis, the pheromones produced by the invading nucleus diffuse or are transported ahead and act as advance signals of nuclear migration. This activates receptors encoded by resident nuclei in nearby cells and the interactions prepare the cells for nuclear migration by initiating septum dissolution to allow nuclei to pass through. A-factor functions then establish the dikaryotic state and clamp connection formation. As mentioned above, pcc1 and clp1, also regulate clamp connection formation. In the haploid (homokaryon), the A-regulated developmental pathway leading to clamp formation is normally suppressed by the pcc1 transcription factor. The HD1/HD2 heterodimer formed by a compatible mating, activates the expression of clp1, which in turn releases the pcc1 repression. A single-gene mutation in pcc1 leads to a complete program of sexual development, including fruit-body morphogenesis and meiosis, suggesting that the pcc1 gene plays a pivotal role in sexual development in Coprinopsis cinerea (Kamada, 2002).
The A and B mating type pathways in Coprinopsis cinerea monokaryons can be activated by transformation with cloned genes from strains of compatible mating types (Kües, Walser, Klaus & Aebi, 2002) and this strategy has been used to show that the A mating type genes regulate production of submerged chlamydospores, hyphal knots and sclerotia in cultures kept in the dark. With illumination, fruit body primordia developed but these aborted before karyogamy (also, illumination permits production of oidia, which is repressed by the action of A mating type genes in the dark). B mating type genes enhance the effects of the A genes on developmental processes, and partially repress the negative action of light on A-mediated regulation of development. Activation of the B pathway alone is not sufficient to induce fruiting, but A-induced fruiting occurs more efficiently and earlier when the B mating type pathway is also active and A+B-activated transformants of the experimental monokaryon formed mature fruit bodies. The B genes are concluded to control fruit body maturation from karyogamy onwards (Kües, Walser, Klaus & Aebi, 2002). It has been suggested that A mating type genes and blue light perception are involved in control of all major differentiation processes in Coprinopsis (Kües, Granado, Hermann, Boulianne, Kertesz-Chaloupková & Aebi, 1998) (discussed in section 2 Stem and cap morphogenesis in Coprinopsis fruit bodies [VIEW NOW]).
Although protein kinases do not feature in the discussions of the last five paragraphs, in animal systems protein kinases are key regulators of cell function. Through the addition of phosphate groups to substrate proteins, they direct the activity, localisation and overall functioning of many proteins, and serve to organise the activity of almost all cellular processes. Kinases are known to regulate the majority of cellular pathways, especially those involved in signal transduction and co-ordination of complex functions like the cell cycle. It must, therefore, be significant that the molecular annotation of the chromosomes of Coprinopsis cinerea completed by Stajich, Wilke, Ahren et al. (2010) revealed so many protein kinase genes. These authors comment (in their supplementary information):
‘The FunK1 family members are of particular interest, because these kinases appear to be restricted to multicellular fungi. Differential transcription of several of these family members has been noted during specific steps of dikaryon formation ... To date, differential transcription of the two FunK1 family members that lie within the B locus … has not been detected. … C. cinerea has 16 members of the tyrosine kinase-like (TKL) kinase group, which are the likely progenitor of animal tyrosine kinases. This group of kinases is found in the Basidiomycetes but is entirely absent from the Saccharomycotina. In plants and Dictyostelium discoideum [the cellular slime mould], TKLs often function as receptor kinases, but no fungal members appear to be membrane linked. In fact, no conventional or atypical C. cinerea kinase is predicted to contain a transmembrane helix domain; if any of these kinases transduce intercellular signals, they presumably do so through a distinct set of nonkinase receptors.’ (Stajich, Wilke, Ahren et al., 2010).
The FunK1 family is the largest kinase family in C. cinerea, having 133 members. This family is (so far in this genomic analysis) specific to Agaricomycotina and Pezizomycotina, but has not been found in other fungi, suggesting a potential link between this kinase family and the multicellularity of the fungi in which it occurs. Unfortunately, it is not at all clear yet how this kinase family is concerned with multicellularity. Taylor & Ellison (2010) put it this way:
‘In C. cinerea, there are 380 genes with protein kinase domains, including many not found in yeast and some previously found only in animals. The largest such family contains the FunK1 domain, which is found in complex Basidiomycota and Ascomycota and is absent in other fungi. Almost one half, 59, of the proteins with this domain are packed into one subtelomeric region of one chromosome. A reasonable hypothesis is that FunK1 domains are important to the development of complex, multicellular fungi and that selection has been involved in retaining these genes over numerous, tandem gene duplications. Other similar and equally reasonable hypotheses could be framed for the P450 and hydrophobin genes. However, despite significant advances related to the mechanisms of genome evolution that emerged from this analysis of the C. cinerea genome, the goal of understanding, with confidence, the genomic basis of developmental complexity remains elusive.’ Taylor & Ellison (2010).
Given the prevalence of data which indicate that hyphal systems (a) need to develop a state of competence before they are able to undertake a developmental pathway, and (b) can be precipitated into embarking upon a particular morphogenesis by a variety of environmental signals, it is difficult to believe that fungi lack developmental management sequences. The only strategies that have been suggested so far that have the potential to be widely used as regulators throughout the higher fungi, are translational triggering and feedback fixation (Timberlake, 1993).
Translational triggering. Translational triggering is a mechanism which can relate a morphogenetic pathway to the development of competence on the one hand, and to initiation in response to environmental cue(s) on the other hand. There are indications from a wide range of physiological studies that nitrogen metabolism may be crucial in regulating morphogenesis. There would certainly be scope for associating particular differentiation pathways with particular aspects of metabolism, so that supply of specific aminoacyl-tRNA molecules might regulate entry into differentiation pathways by affecting translation of a controlling reading frame (trigger-ORF in Fig. 46).
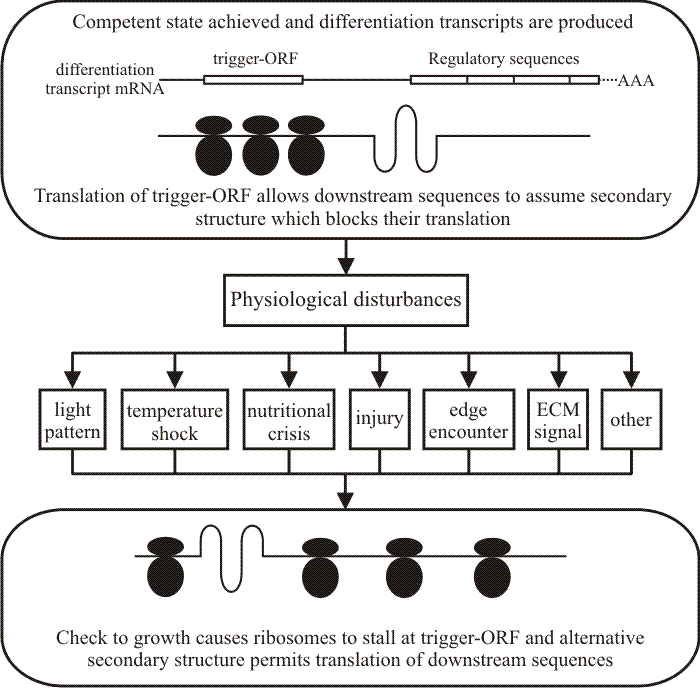 |
| Fig. 46. Translational triggering adopted as a general model for entry of competent tissues into fungal pathways of differentiation. Modified from Moore, 1998a. |
If the trigger-ORF contained adjacent codons for aminoacyl-tRNAs subject to variation in supply, stalling/non-stalling of translation of trigger-ORF might determine whether the messenger transcript forms secondary structures which permit/do not permit translation of down-stream reading frames. Note that either one or both components of the aminoacyl-tRNA may be the limiting factor and the limitation may be imposed by a compartmentalisation. That is, amino acid or a specific tRNA (or, presumably, an aminoacyl-tRNA synthetase) may be compartmentalised, regulated in local concentration, or both.
The interpretation offers a way by which a competent tissue can be released to undertake differentiation by a range of physiological events. Competence is interpreted to mean that messenger transcripts for the necessary regulators (and perhaps some key structural genes) are produced but not fully translated because an upstream sequence (trigger-ORF) prevents translation. There may be a number of different such transcripts with regulators corresponding to the different pathways upon which the competent cell can embark, their trigger-ORFs responding to separate physiological events. On the other hand, there may be a number of similar transcripts in different cellular compartments so that the translational trigger can be released by the particular activities of those compartments with the result that one differentiation process may be triggered by different physiological events. It could also be that such a transcript was limited to one compartment, even one type of vesicle, perhaps, from which the trigger molecule can be excluded until some highly specific and/or localised physiological change occurs.
Unfortunately, there is no evidence for any of these speculations. The only possibly relevant data is the finding that depletion of carbon source available to Saccharomyces cerevisiae blocks translation but not transcription (Martinez-Pastor & Estruch, 1996). This may not even be relevant, but it does at least indicate that a major physiological crisis can have more immediate impact on translation and it emphasises that carbon metabolism is important, too, even if nitrogen metabolism is more often associated with morphogenetic change. Despite the lack of direct evidence at this time, I suggest that a variety of physiological signals and stresses cause translation-level controls to direct competent fungal tissues to undertake specific differentiation processes. Comparison with the operation of mating type factors makes it reasonable to suggest that the translational trigger could immediately lead to translation of components of highly specific transcription activators and inhibitors which then regulate gene sequences required for the differentiation which has been initiated. These, or their eventual products, may be involved in feedback fixation of the differentiation pathway.
Feedback fixation. Feedback fixation is the outcome of feedback activation and autoregulation which together reinforce expression of the whole regulatory pathway to make it independent of the external environmental cues which initiated it. Feedback fixation results in developmental determination in the classic embryological sense. The epigenetic aspect of the network governing fungal morphogenesis starts with feedback fixation, but also includes signals from outside the cell (Fig. 47). The fungal extracellular matrix is extensive and complex. Its reaction to, and interaction with the environment can be communicated to the intracellular environment to modify cytoplasmic activity. Since neighbouring cells are components of the external environment, it must be the case that the activity of one hyphal cell is modulated by changes made to the extracellular matrix by a neighbouring hyphal cell.
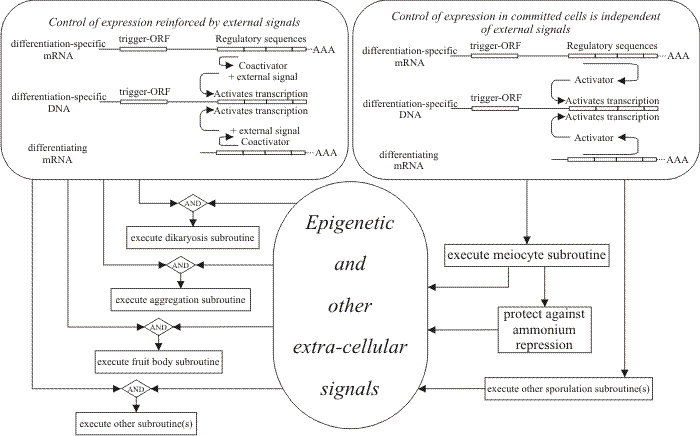 |
| Fig. 47. Feedback fixation adopted as a general model for maintaining progress through fungal pathways of differentiation. In this flow chart the boxes at the top show, on the left, the type of feedback fixation process envisaged to apply to most developmental subroutines in which epigenetic reinforcement from the local microenvironment is needed to interact with coactivators in order to maintain the feedback activation loop. In the panel at top right, the alternative of direct feedback fixation independently of other signals is shown as being applicable to cell types which show developmental commitment; only meiocytes are known to be committed, but there may be other committed cell types. Revised from Moore, 1998a. |
On this interpretation, therefore, continued progress in differentiation for most fungal cells requires continued reinforcement from their local microenvironment. This may involve production of location- and/or time-specific extracellular matrix molecules, or any of a range of smaller molecules which might be classed as hormones or growth factors. Smaller molecules might exert their effects by being taken up into the cell. But uptake is not necessary. Any of these molecules may also affect relations between integrins and the existing extracellular matrix. As a result there could be direct effects on the cytoskeleton which are able to cause immediate metabolic changes in one or more cellular compartments, or directly influence gene transcription.
Connections to the extracellular matrix may also be involved in that other great enigma: the control of hyphal branching. By varying extracellular matrix/membrane or wall/membrane connections external signals may be able to specify branch initiation sites. Similarly, internal cytoskeletal architecture could also arrange specific membrane/wall connections to become branch initiation sites. Branch initiation sites specified in these ways may then become gathering sites for the molecules which create a new hyphal tip. First among these could be molecular chaperones (heat shock proteins; Hartl, Hlodan & Langer, 1994) which rearrange existing wall and membrane proteins. The chaperones could arrange for a mature wall region to become juvenile again by altering the conformation of existing polypeptides rather than by severing any covalent bonds. Local activation of resident chitin synthase (Gooday & Schofield, 1995) and initiation of a localised resumption of wall synthesis would then establish the new hyphal tip. The branch would consequently emerge in a position precisely defined by the stimulation of generalised cytoskeleton/membrane/wall connections by a positional stimulus.
The focus of these hypothetical regulatory activities is, obviously, the hyphal wall, its surface and the immediate extracellular environment. These are features about which we are very ignorant and urgent and extensive research on these topics is necessary. The key to fungal morphogenesis lies in understanding how that which is outside a hypha can influence that which goes on inside the hypha in a time- and place-dependent manner. We are still a long way from reaching that understanding. [TOP]
Updated December 7, 2016










