10.12 Primary (intermediary) metabolism
Metabolic pathways which characteristically operate when a fungus is growing at or near its maximum rate are described as primary pathways; secondary pathways become operational (or amplified) when growth rate is limited in some way to a level below the maximum (Bu’Lock, 1967). The fundamental function of primary metabolism is the utilisation of nutrients to form ATP and reduced nucleotide coenzymes (NADH and NADPH), which together equal chemical energy and reducing power, and the compounds which serve as precursors of cellular components, especially macromolecular constituents. All living organisms adapt continuously to changes in their environment and particularly to variation in the availability of energy-yielding substrates. Extensive studies have identified protein acetylation as an important mechanism to regulate metabolic enzymes by activation, inhibition, by influencing protein stability, and by regulation of transcription. Proteomic analyses reveal many acetylated proteins in the cytoplasm and mitochondria, including most enzymes involved in intermediate metabolism (Guan & Xiong, 2011). Numerous links between the products of intermediary metabolism and the cell’s transcriptional machinery form a complex network that integrates environmental inputs to produce the most appropriate responses from the genome (Gut & Verdin, 2013).
The major source of energy and reducing power is the catabolism of carbohydrate. Although other carbon-containing compounds can be utilised for these purposes by most cells, the full sequence of enzymic processes are conventionally represented as involving the controlled release of energy by the use of atmospheric oxygen to convert glucose to CO2 and water. This overall process is described as respiration and its chemically balanced (or stoichiometric) summary equation is:
C6H12O6 + 6O2 → 6CO2 + 6H2O + energy
Note that this equation does not even begin to describe the biochemical mechanisms which achieve the indicated chemical transformation, but it does emphasise that for the conversion of each mole of glucose, six moles of oxygen must be absorbed from the atmosphere, and six moles of CO2 and six moles of water appear within the cell, and 2 900 kJ of free energy (i.e. energy capable of doing some work) are released. To put this into more readily grasped units, respiration of 1 g of glucose uses 1.07 g (747 cm3) oxygen and produces 1.47 g (747 cm3) CO2 and 0.6 g water, releasing 16.1 kJ of energy.
To achieve this basic chemistry the living cell uses a sequence of enzymically controlled reactions. These are conveniently divided into three phases or subpathways:
- glycolysis including the pentose phosphate pathway (PPP),
- the tricarboxylic acid (TCA, or Krebs) cycle,
- oxidative phosphorylation.
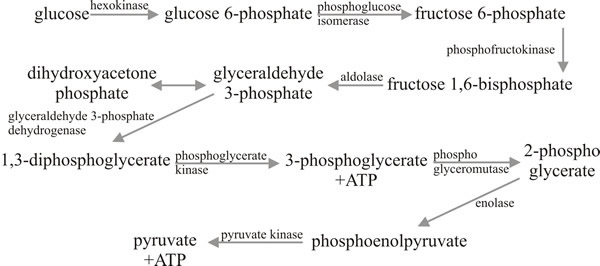 |
Fig. 7. The Embden-Meyerhof-Parnass (EMP) glycolytic
pathway. |
The word glycolysis describes the conversion of glucose to pyruvate without implying a particular pathway. In fact, there are three enzymic pathways which might be used, though one does tend to predominate. The Embden-Meyerhof-Parnass (EMP) pathway is the major pathway in most species; it comprises nine enzymic steps, all of which occur in the cytoplasm (Fig. 7). The net outcome of the reactions summarised in Fig. 7 is that one molecule of glucose is converted to two molecules of pyruvic acid plus two molecules of ATP and two molecules of NADH2. Thus, the energy yield (2ATP + 2NADH2, since the latter do represent potential chemical work), is rather small; the main function of the EMP pathway being conversion of glucose to pyruvate for processing in the TCA cycle.
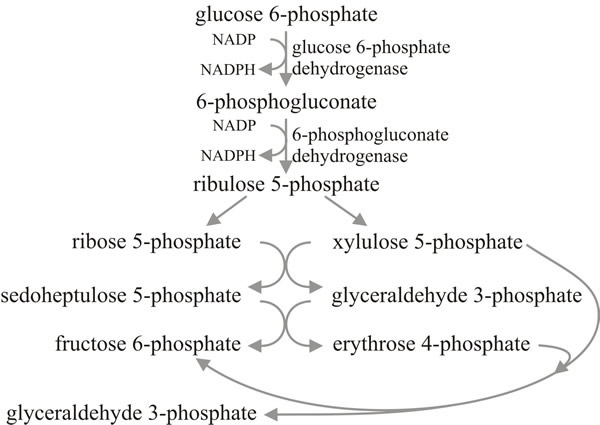
A commonly encountered alternative glycolytic pathway is the pentose phosphate pathway (PPP) (Fig. 8). This is also called the hexose monophosphate pathway (HMP) and in strictly chemical terms such a name is accurate; glucose 6-phosphate is diverted out, undergoes a range of chemical conversions, and then fructose 6-phosphate and glyceraldehyde 3-phosphate feed back into the EMP pathway. But the name pentose phosphate pathway does acknowledge that the PPP provides pentose sugars for nucleotide synthesis (which includes coenzymes and energy carriers as well as RNA and DNA), erythrose phosphate for the synthesis of aromatic amino acids through the shikimic acid pathway, and NADPH2, which is the coenzyme most often used in biosynthetic reactions that require reducing power, especially fat and oil synthesis. So although the PPP can theoretically achieve complete glycolysis (six cycles through the reaction sequence would completely oxidise a molecule of glucose to CO2), it is more likely to be involved in furnishing biosynthetic intermediates. The PPP also, of course, provides a route for utilisation of pentose sugars which become available as carbon sources, and for interconverting hexose and pentose phosphates.
 |
There is a third glycolytic pathway, the Entner-Doudoroff (ED) pathway (not illustrated here) that proceeds via 6-phosphogluconate to 2-keto-3-deoxy-6-phosphogluconate, which gives rise to pyruvate and glyceraldehyde 3-phosphate. It is a common glycolytic pathway in bacteria, but has been demonstrated in only a few fungi.
The use of different glycolytic pathways in any cell will reflect the relative contribution their intermediates are required to make to the functions of the cell; they will change with age, activity and nutrition. In general, since the PPP provides intermediates for biosynthesis, use of this pathway increases in rapidly growing and in differentiating cells, and is minimised in those which are resting or quiescent.
Whichever glycolytic pathway contributes the pyruvate, this latter molecule is formed in the cytoplasm and must then be transported into the mitochondrion where it is converted to acetyl coenzyme A (acetyl-CoA). This step is achieved by the pyruvate dehydrogenase complex; a combination of enzymes which first decarboxylate pyruvate and then transfer the resulting acetyl group to coenzyme A. The pyruvate dehydrogenase complex is emerging as a major control point of oxidative phosphorylation in mitochondria (Acin-Perez et al., 2010).
The TCA cycle is cyclic because its ‘end-product’, oxaloacetate, reacts with acetyl-CoA to introduce what remain of the pyruvate carbon atoms into a reaction sequence (Fig. 9) the primary function of which is to convert pyruvate formed in glycolysis entirely to CO2, the released energy being captured primarily in NADH2. The overall stoichiometry is that one molecule of pyruvate with three molecules of water forms three molecules of CO2 and releases 10 protons, the latter appearing in the form of three molecules of NADH2, and one each of FADH2 and the ‘high energy’ compound GTP. The succinate dehydrogenase enzyme is bound to the inner mitochondrial membrane (and, because of this, is often used as a marker for the presence of mitochondria in fractionated cell extracts), the other enzymes of the TCA cycle occur in the mitochondrial matrix.
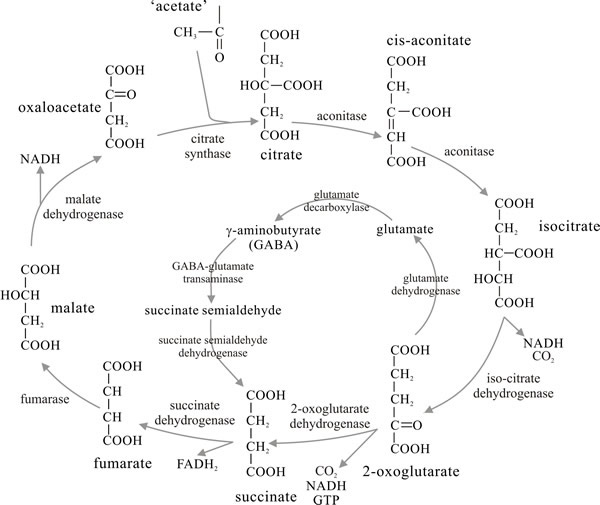 |
A common variant of the TCA cycle is the glutamate decarboxylation loop in which 2-oxoglutarate is aminated to glutamate rather than being oxidatively decarboxylated to succinate. The glutamate is decarboxylated to 4-aminobutyrate; transamination between the latter and 2-oxoglutarate yielding succinate semialdehyde which, on oxidation, feeds back into the TCA cycle as succinate (central panel in Fig. 9). The enzymes of this loop have been found to be at high activity in fruit bodies, especially caps, of the ink cap mushroom Coprinopsis cinerea, where it is the normal route of TCA metabolism in this organism. The glutamate decarboxylation loop also operates in Agaricus bisporus.
Through glycolysis and the TCA cycle, all of the carbon contained in the substrate glucose is released as CO2. However, very little energy is released, most of it being captured in NADH2 (with a small amount in GTP and FADH2). The energy represented in the form of the reduced coenzymes is recovered as ATP through the electron transport chain, located on the inner mitochondrial membrane, in the process known as oxidative phosphorylation. The electron transport chain transfers electrons from the reduced coenzymes through a series of reactions until the electrons are finally passed to oxygen, reducing it to water. Stepwise transfer of electrons between components of the electron transport chain leads to the pumping of protons from the mitochondrial matrix into the intermembrane space.
The resulting proton gradient (the pH in the intermembrane space is about 1.4 pH units lower than that of the matrix) is used to generate ATP. Transfer of a pair of electrons from one molecule of NADH2 to oxygen leads to proton pumping at three sites in the chain, at each of which the consequent proton gradient can be used to synthesise one molecule of ATP. The ATP is synthesised by an enzyme complex located on the matrix side of the inner mitochondrial membrane. As protons move down a channel in this complex (the channel, known as the F0 sector, is composed of at least four hydrophobic subunits forming the proton channel located in the membrane) the associated F1 sector (containing five different subunits) projects into the matrix and is responsible for ATP synthesis (see Chapter 5) [to view an animation describing ATP synthase powered by a proton gradient visit http://vcell.ndsu.nodak.edu/animations/atpgradient/index.htm or watch the video on YouTube at: https://www.youtube.com/watch?v=3y1dO4nNaKY]).
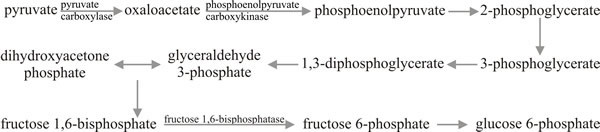 |
Fig. 10. Gluconeogenesis: the making of carbohydrates.
Modified from Moore, 1998. |
Although we are primarily concerned here with catabolism, the pathways described permit sugar synthesis with just a few modifications. We stressed above that glycolysis and the TCA cycle provide opportunities for the fungus to make use of a very wide range of potential carbon and energy sources; but one which is successfully growing on acetate, for example, is clearly required to synthesise all of those compounds which have more than two carbon atoms chained together. In such circumstances glycolysis cannot simply be reversed because the steps governed by kinases (hexokinase, phosphofructokinase and pyruvate kinase) are irreversible, so for these steps in particular, additional enzymes are required for gluconeogenesis (Fig. 10).
In these circumstances the first steps in conversion of pyruvate to carbohydrate are carried out by pyruvate carboxylase, which synthesises oxaloacetate which is then decarboxylated and phosphorylated to phosphoenolpyruvate by phosphoenolpyruvate carboxykinase. The phosphoenolpyruvate can then be converted to fructose 1,6-bisphosphate by reversal of the EMP pathway, but an additional enzyme, fructose bisphosphatase, is required to generate fructose 6-phosphate. As the sugar phosphates are readily interconvertible, once this compound is formed oligosaccharide and polysaccharide synthesis can proceed. The structures of many of the polysaccharides formed have been shown earlier in this chapter.
Glycolysis and gluconeogenesis are obviously alternatives which demand close control to assure metabolic balance. Phosphofructokinase is the key glycolytic control point, and fructose bisphosphatase responds inversely to the same molecules, being allosterically activated by citrate but inhibited by AMP.
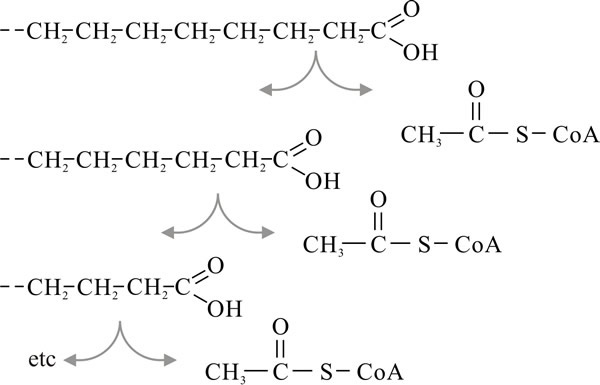 |
Fig. 11. β-oxidation of fatty acids. Modified from
Moore, 1998. |
Before leaving carbohydrate metabolism, it is worth mentioning here that the sugar alcohol mannitol and the disaccharide trehalose are almost always found among the water-soluble cytoplasmic carbohydrates in fungi; trehalose being the most widely-distributed sugar in fungi. Mannitol and trehalose seem to serve as transient storage compounds (i.e. molecules capable of immediate mobilisation when required) and both have been identified as substrates used for the metabolism associated with spore germination.
Trehalose synthesis/accumulation/degradation cycles occur at a number of stages in development of a fungus so it is quite clear that this sugar is the ‘common currency’ of the fungal carbohydrate economy. It is synthesised, as trehalose 6-phosphate, by the enzyme trehalose phosphate synthase from glucose 6-phosphate and the sugar nucleotide UDP-glucose. There may be other functions for mannitol. It can certainly serve an osmoregulatory function in the marine fungus Dendryphiella and may serve the same purpose in fruit bodies of the cultivated mushroom, Agaricus bisporus, in which it can be accumulated to concentrations of up to 50% of the total dry weight (yes, that’s right, half of the cultivated mushroom on your plate is mannitol); similar concentrations have been encountered in fruit bodies of Lentinula edodes (shiitake). In A. bisporus, mannitol is synthesised by reduction of fructose by an NADP-linked mannitol dehydrogenase.
Fats are molecules of glycerol in which the three hydroxyl groups are replaced with three fatty acid molecules. In degradation the first step is carried out by lipase which removes the fatty acids from the glycerol. The latter can be converted to glyceraldehyde 3-phosphate and thereby enter glycolysis (see Fig. 8), but it represents only about 10% by weight of a fat molecule, the bulk being represented by the fatty acids which consist of long carbon chains (e.g. palmitic acid, C16; stearic acid, C18).
These chains are degraded by sequential removal of the two terminal carbon atoms in the form of an acetyl group attached to CoA (Fig. 11). Because the cleavage occurs at the second (β) carbon atom, this process is called β-oxidation and it takes place in the mitochondrial matrix. Each such cleavage is oxidative, enzymes passing the H-atoms to the coenzymes NAD and FAD. Thus, degradation of palmitic acid requires seven cleavages and yields eight molecules of acetyl-CoA (which enter the TCA cycle), seven NADH2 and seven FADH2 (both of which enter the electron transport chain for oxidative phosphorylation). Oxidation of fatty acids releases considerable amounts of energy; for example, one molecule of palmitic acid will give rise to about 100 molecules of ATP. This is why fats are such effective energy storage compounds.
Ultimately, all nitrogen in living organisms is derived from the native element in the atmosphere. Each year an amount between 100 and 200 million tonnes of atmospheric nitrogen is reduced to ammonium by the nitrogenase enzyme system of nitrogen-fixing bacteria and blue-green algae. On the basis of present knowledge, it is unlikely that any fungi are able to fix elemental nitrogen.
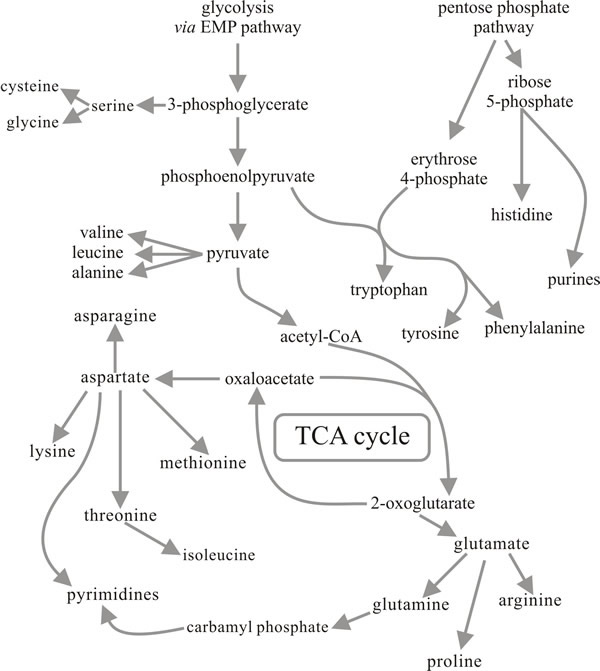 |
If a fungus is unable to access amino groups by direct absorption of amino acids, they have to be formed and the most immediate source is by the assimilation of ammonium. The only route of ammonium assimilation which can be considered pretty well universal in fungi is the synthesis of glutamate from ammonium and 2-oxoglutarate by the enzyme glutamate dehydrogenase. Many filamentous fungi and yeasts have been shown to produce two glutamate dehydrogenase enzymes; one linked to the coenzyme NAD and the other linked to NADP.
As can be appreciated from Fig.12, the interconversion of 2-oxoglutarate and glutamate is a reaction which occupies a central position in metabolism and is one at which important pathways in both carbon metabolism and nitrogen metabolism come together. The reaction is readily reversible and it is often considered that NAD-linked glutamate dehydrogenase (NAD-GDH) has a deaminating or catabolic role (glutamate → 2-oxoglutarate + ammonium), while the NADP-linked enzyme provides the aminating or anabolic function (2-oxoglutarate + ammonium → glutamate). However, some organisms have evolved different patterns of endogenous regulation, especially in relation to their morphogenetic processes. For example, in the ink cap mushroom Coprinopsis cinerea the NADP-GDH normally appears at high activity only in the cap tissue of the mushroom fruit body where it is located in the basidia apparently protecting meiosis and sporulation from inhibition by ammonium, i.e. acting as an ammonium detoxifier rather than ammonium assimilator.
This example aside, NADP-GDH is generally the most important enzyme involved in ammonium assimilation in mycelia and its activity is often increased when ammonium is provided as a growth-limiting, sole nitrogen source. However, in some fungi an alternative enzyme system appears to scavenge for ammonium when it becomes limiting, this is the glutamine synthetase/glutamate synthase system. Glutamine synthetase is widely, perhaps universally, distributed and is responsible for synthesis of glutamine. However, glutamine synthetase can have a high affinity for ammonium and, in combination with glutamate synthase (which converts glutamine + 2-oxoglutarate to two molecules of glutamate), forms an ammonium assimilation system which can recover ammonium even when this molecule is present at extremely low concentrations. The net result (2-oxoglutarate + NH4+ → glutamate) is the same as the reaction promoted by NADP-GDH, but the cost is higher as glutamine synthetase uses ATP to make glutamine. The glutamate synthase mechanism is common in bacteria and is encountered in fewer fungi, but has been demonstrated in Neurospora crassa, Aspergillus nidulans and several yeasts and mycorrhizal fungi.
Some yeasts and a larger number of filamentous fungi can utilise nitrate as sole source of nitrogen. Chemically, nitrate is first converted to nitrite which is then converted to ammonium, but the enzymic steps are quite complicated. The complexity of the reaction is reflected in the large number of mutant genes, in both Neurospora and Aspergillus, which have been found to affect nitrate assimilation. The first stage is performed by nitrate reductase which, generally in fungi, has a cofactor containing molybdenum and requires NADPH (NADH in at least some yeasts). Nitrate is thought to bind to the molybdenum-cofactor of the enzyme prior to being reduced by removal of an oxygen atom. Removal of this allows the nitrate formed to be bound to nitrite reductase, through its nitrogen atom, for the reduction to ammonium. The ammonium formed from nitrate is immediately used for the reductive amination of 2-oxoglutarate to glutamate.
Conversion of nitrate (NO3-) to ammonia (NH3) is a chemical reduction requiring considerable energy expenditure. In fact the equivalent of four NADPH2 molecules (880 kJ of energy) are used to reduce one NO3- ion to NH3 , which is additional to the energy demand for assimilation of the ammonium (one NADPH2 is used for assimilation via NADP-GDH, 1 NADPH2 + 1 ATP for assimilation through glutamine synthetase and glutamate synthase). Given these additional energy demands, it is not surprising that the nitrate reduction machinery is produced only when nitrate is the sole available source of nitrogen, being induced by nitrate and rapidly repressed by the presence in the medium of ammonium or any alternative sources of reduced nitrogen.
The constituents of living cells are in a continual state of flux; all components being subjected to turnover as old materials are catabolised and new ones synthesised. When proteins and other nitrogen-containing compounds are broken down, either as part of this turnover process or as externally supplied nutrients, the carbon can be disposed of as CO2, hydrogen as water and nitrogen either as ammonium or as urea. The use of protein as a carbon source has been discussed above. In these circumstances the organism (animal, plant or fungus) suffers an excess of nitrogen and must excrete it. Experiments with the basidiomycetes Agaricus bisporus, Coprinopsis cinerea and Volvariella volvacea have shown that one third to one half of the nitrogen contained in the protein given as substrate is excreted as ammonium into the medium.
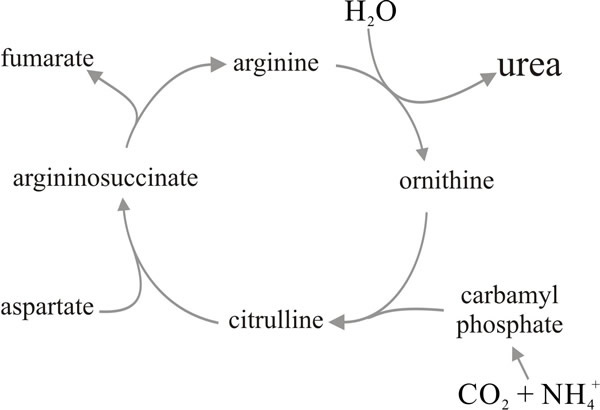
In terrestrial mammals metabolising protein the toxicity of ammonium is avoided by excretion of urea formed through the urea cycle (Fig. 13). However, the enzyme urease seems generally to be constitutive in fungal mycelia so any urea formed is likely to be dissimilated to NH3 and CO2. Nevertheless there are circumstances in which fungi accumulate urea (at which time they repress the urease). Especially large accumulations have been found in fruit bodies of Basidiomycota, where it seems likely that it acts as an ‘osmotic metabolite’ controlling water entry into cells during expansion of the fruit body tissues. Thus, the capacity to synthesise, and even accumulate, urea is well developed in fungi but it seems that it is ammonia which is excreted to dispose of excess nitrogen.
 |
