10.7 Lignin degradation
Lignins are high-molecular weight, insoluble aromatic polymers present predominantly in the walls of secondarily thickened plant cells, making them rigid and impervious. They are composed essentially of many methoxylated derivatives of benzene (phenylpropanoid alcohols, also called monolignols), especially coniferyl, sinapyl, and coumaryl alcohols (Fig. 5), the proportions of these three differ between angiosperms and gymnosperms, and between different plants. Lignins are highly heterogeneous polymers, the monolignols being assembled to suit the lignin to different functions in the plant cell wall (Humphreys & Chapple, 2002; Davin & Lewis, 2005; Vanholme et al., 2010; Shafrin et al., 2017).
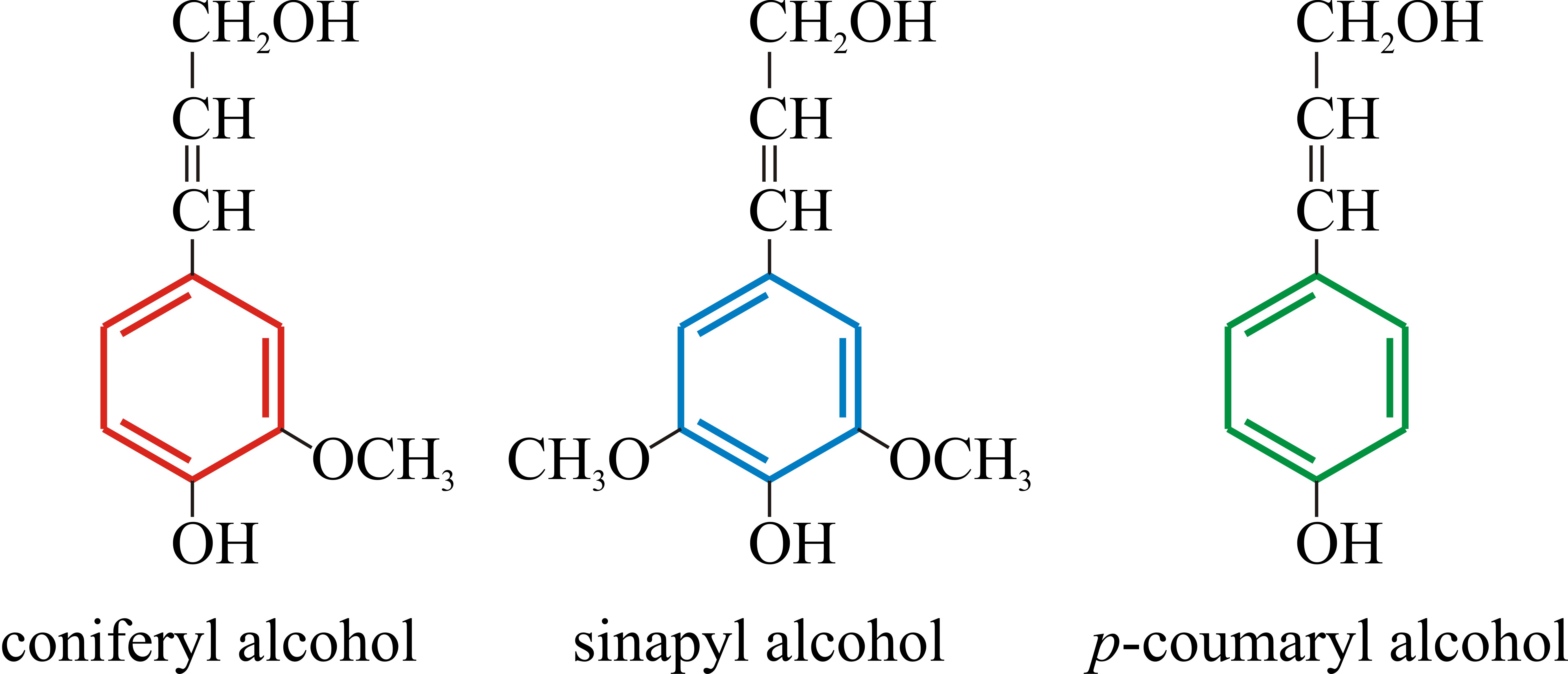 |
Fig. 10.5. Chemical structures of the phenylpropanoid
alcohols used to construct the lignin polymer. These are also called
monolignols. They are colour coded here to indicate how they probably
contribute to the lignin polymer illustrated in Fig. 10.6. The
proportions of these three phenylpropanoid alcohols differ between the
lignins of angiosperms and gymnosperms, and between lignins of different
plant species. |
You are not expected to ‘know’ the primary structure of angiosperm lignin (Fig. 6) in the sense of being able to draw it out yourself, but you are expected to understand what it is telling you about the nature of lignin as a nutritional substrate and the enzymological challenge it poses to any organism setting out to degrade it. Note the way the three phenylpropanoid alcohols are used in the structure; note the predominance of ether linkages (-C-O-C-) and carbon-carbon bonding, and the presence of a few hydroxyls that can take part in cross links to other polymers (polysaccharides and proteins, for example). Above all, note the predominance of benzene rings. How many foods do you know that are full of benzene rings and how would you metabolise a benzene ring? In your kitchen, benzene rings, especially phenols, are more likely to be found in disinfectants than foods, and that’s the key to the function of lignin. Micro-organisms that attempt to degrade lignin produce anti-microbial disinfectants, which makes lignin very resistant to attack.
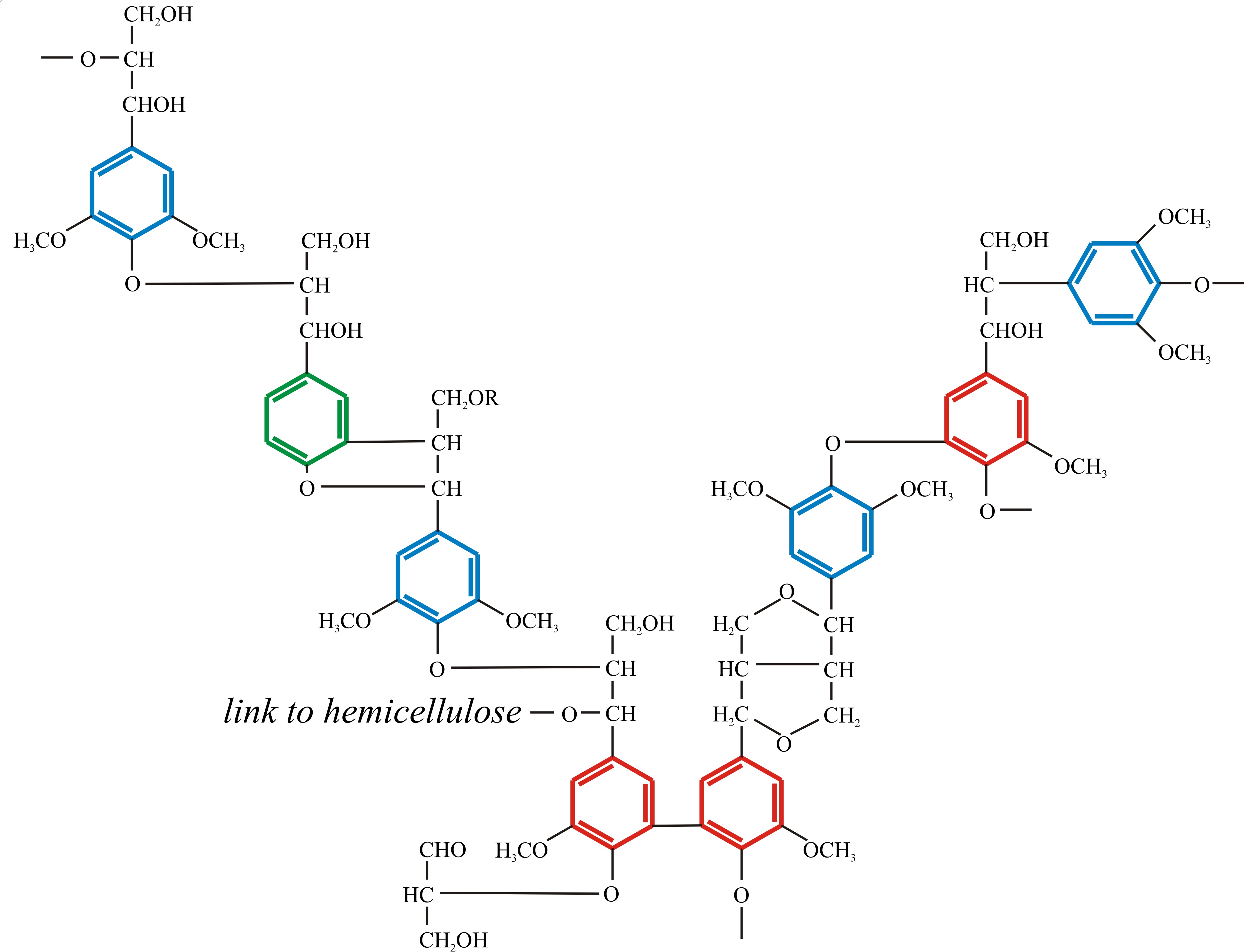 |
Fig. 10.6. Schematic formula of the polymer structure
of angiosperm lignin. Note the way the three phenylpropanoid alcohols
are used in the structure (colour coded to match those in Fig. 10.5);
note the predominance of ether linkages and carbon-carbon bonding, and
the presence of a few hydroxyls that can take part in cross links to
other polymers (polysaccharides like hemicellulose and proteins). Above
all, note the predominance of benzene rings. Modified from Moore, 1998;
and see Vanholme et al. (2010) for more structural details. |
This extreme resistance to microbial degradation is part of the principal function of lignin as it can protect other polymers from attack. We are used to hydrolases cleaving polymers (polysaccharides, proteins, lipids), but the carbon-carbon and ether bonds that join subunits together in lignin must be cleaved by an oxidative process and a range of enzymes are needed for lignin to be degraded. Although ability to digest simple synthetic lignins has been reported for a few bacteria, ability to degrade natural lignins is generally considered to be limited to a very few fungi. The list includes a range of Basidiomycota and Ascomycota, and the fungi involved are generally known as ‘white-rots’ because the lignin they digest away provides the main pigmentation of wood.
Oxidative lignin breakdown depends on two major groups of enzymes called heme peroxidases and laccases. Lignin-degrading peroxidases include:
- lignin peroxidase (LiP; a heme [Fe]-containing protein) which catalyses H2O2-dependent oxidation of lignin). LiPs are strong oxidants that oxidise the major non-phenolic structures of lignin;
- manganese-dependent peroxidase (MnP, which also catalyses H2O2-dependent oxidation of lignin). MnP catalyses the oxidation of various phenolic substrates but is not capable of oxidising the more recalcitrant non-phenolic lignin;
- versatile peroxidase (VP; also called manganese-independent peroxidase, another heme-containing peroxidase with the catalytic activities of both MnP and LiP). It can oxidise Mn2+, like MnP, but also non-phenolic compounds, like LiP (VP has only been described in species of the genera Pleurotus and Bjerkandera);
- dye-decolourising peroxidase (DyP; which uses H2O2 to decolourise anthraquinone dyes). DyPs occur in both fungi and bacteria.
LiP, MnP, and VP are class II extracellular fungal peroxidases that belong to the plant and microbial peroxidases superfamily. DyPs are members of a new superfamily of heme peroxidases reported to oxidise lignin model compounds.
The second major group of lignin-degrading enzymes, laccases, are copper-containing proteins found in plants, fungi, and bacteria. They belong to the multicopper oxidase superfamily and are described in a little more detail below. Fungal laccases catalyse demethylation of lignin components and can oxidise phenolic lignin model compounds; they have higher redox potential than bacterial laccases. In the presence of redox mediators, fungal laccases can also oxidise non-phenolic lignin model compounds (a redox mediator is a molecule that is stable in two interconvertible redox states, and is sufficiently small to diffuse easily into and out of protein channels and efficiently shuttle electrons between the enzyme active site and the substrate).
In addition to the peroxidases and laccases, fungi produce other accessory oxidases such as aryl-alcohol oxidase, also called veratryl alcohol oxidase (veratryl alcohol is a degradation product of lignin) and glyoxal oxidase that generate the hydrogen peroxide required by the peroxidases (Sigoillot et al., 2012; Abdel-Hamid et al., 2013).
The process of catabolic lignin degradation involves:
- cleavage of ether bonds between monomers;
- oxidative cleavage of the propane side chain;
- demethylation;
- benzene ring cleavage to ketoadipic acid which is fed into the tricarboxylic acid cycle as a fatty acid.
Most research has been concentrated on white-rot basidiomycete fungi, such as Phanerochaete chrysosporium (= Sporotrichum pulverulentum), which is able to mineralise lignin completely to carbon dioxide and water. The lignin-degradative system of P. chrysosporium appears after cessation of primary growth (that is, it is an aspect of the secondary metabolism of the organism) and can be induced by nitrogen starvation. Genomes of white-rot fungi feature families of genes encoding laccases, lignin peroxidases and manganese peroxidases. The multiplicity of these extracellular enzymes could be a response to the diversity of the lignin substrate but may also be a result of their having multiple roles in fungal physiology (laccases, for example, contribute to plant pathogenesis, sporulation and pigment formation). Specialisation among secreted fungal oxidoreductases (laccases and peroxidases) is indicated by the fact that fungi that favour wood that is already partially degraded (called secondary colonisers) produce a distinctly different spectrum of enzymes than the primary colonisers of wood, and different again from litter decomposers or humus saprotrophs (Moiseenko et al., 2019).
Lignin peroxidase (ligninase) is the key lignin-degrading enzyme of white-rot fungi. The P. chrysosporium family of lignin peroxidases comprises extracellular glycosylated heme proteins, which are secreted in response to nitrogen limitation. When the fungus is grown in low-nitrogen medium there is an increase in H2O2 production by cell extracts which correlates with the appearance of ligninolytic activity; experimental destruction of H2O2 by adding the enzyme catalase strongly inhibits lignin breakdown. Activated oxygen derived from H2O2 is involved in degrading lignin, but is held in the active site of a specific extracellular enzyme, the lignin peroxidase. Lignin peroxidases are strongly oxidative; the enzyme is activated by itself being oxidised by H2O2, the initial step involving oxidation by one electron to produce an unstable intermediate which is then able to catalyse oxidation of phenols, aromatic amines, aromatic ethers and polycyclic aromatic hydrocarbons. Veratryl alcohol is a secondary metabolite that stimulates lignin degradation by recycling the lignin peroxidase and protecting the enzyme against inactivation by H2O2.
There can be as many as 15 lignin peroxidase isozymes, ranging in molecular mass from 38,000 to 43,000, the spectrum of isozymes produced depending on culture conditions and strains employed. However, ten lignin peroxidase genes, with a conserved sequence, have been identified in P. chrysosporium and mapped into three different linkage groups. Lignin peroxidases are not found in all white-rot fungi, though. In particular, they seem to be absent from Ceriporiopsis subvermispora, a white-rot basidiomycete that is widely studied for its potential use in the pulp and paper industry (‘biopulping’; Breen & Singleton, 1999) and in which manganese peroxidases are responsible for ligninolysis.
Manganese-dependent peroxidases are another family of extracellular glycosylated heme proteins, which are produced by most white-rot fungi. Like lignin peroxidase, the manganese peroxidases also require H2O2 to function, but the mechanism is very different. The manganese peroxidase system creates low molecular weight oxidising agents that diffuse into the lignin substrate and are able to oxidise phenolic residues in the lignin some distance from the enzyme. Some of these oxidising agents may be peroxidated lipids, but the chief one, from which the enzyme gets its name, is the metal ion manganese; the enzyme uses H2O2 to oxidise extracellular Mn(II) to Mn(III) and this becomes the diffusible oxidant that can degrade lignin at a distance.
The H2O2 which is required by peroxidases is probably generated by glyoxal oxidase, an extracellular enzyme that transfers electrons from low molecular weight aldehydes (e.g. glyoxal and glycoaldehyde) to O2 and so forms H2O2. Aryl alcohol oxidase is another H2O2 generating enzyme; in the process of converting benzyl alcohols to the aldehydes it transfers electrons to O2, generating H2O2. C. subvermispora secretes an oxalate oxidase, which catalyses the degradation of oxalate to carbon dioxide and H2O2 and could be the main provider of H2O2 for manganese peroxidase in this organism.
Although few fungi produce ligninolytic enzymes, a much wider range excrete laccases as extracellular enzymes. These are copper-containing oxygenases which are able to oxidise o- and p-phenols and are required for the metabolism of lignin degradation-products. They are particularly interesting as their appearance or disappearance in fungal cultures has been correlated with sexual and asexual reproduction in a number of cases. Thus, during mycelial growth of the cultivated mushroom, Agaricus bisporus, a large proportion of the compost lignin is degraded and correspondingly high activities of laccase are recorded (this one enzyme can amount to 2% of the total fungal protein). Yet, as the culture forms fruit bodies laccase activity is rapidly lost, initially by inactivation and subsequently by proteolysis; this pattern of behaviour reflecting the changing nutritional demands of fungal mycelia as they process through successive developmental phases and the ability of the mycelium to act on its environment to satisfy those demands.
Laccases have been found in many fungi, including non-ligninolytic members of the Ascomycota, such as Aspergillus and Neurospora, as well as wood-rotting Basidiomycota. Laccases also occur in plants where they contribute to lignin biosynthesis. To react with a broader range of substrates laccases can interact with a low molecular weight co-substrate secreted by the fungus that functions as a diffusible lignin-oxidising agent (redox mediator).
In their study of laccase, Mn-dependent peroxidase, and versatile peroxidase activities in seven wood-rotting fungi digesting wheat straw lignin, Knežević et al. (2013) found that the profiles of the three enzymes varied between species and during the cultivation period. Dichomitus (Bjerkandera) squalens (Western Red Rot, a poroid crust fungus of conifers) was the best lignin degrader (34.1% of lignin digested), and Pleurotus ostreatus and Pleurotus eryngii were the weakest (7.1% and 14.5% of lignin digested, respectively). These two Pleurotus species produced the most laccase, whereas Dichomitus squalens produced the most versatile peroxidase (with the least in Pleurotus ostreatus). Lenzites betulinus (Birch Mazegill) and Fomitopsis pinicola (red belted polypore, a bracket or shelf fungus on fir and hemlock) were the best Mn-dependent peroxidase producers, with Pleurotus ostreatus the weakest; but the laccase isoforms (isoforms being proteins that have similar but not identical amino acid sequences) produced by Pleurotus ostreatus depend on the conditions in which it is grown. The presence of inducers, the pH of the culture medium as well as the type and concentration of carbon and nitrogen sources all differentially affect the production of these enzymes, and the growth temperature also modifies the patterns of activity and production of laccase isoforms (Montalvo et al., 2020).
Lignin-degrading enzymes have considerable promise for several biotechnological applications such as paper industry, textile industry, wastewater treatment and the degradation of herbicides. Biopulping has already been mentioned as an industry where ligninolytic enzymes can improve the quality of pulp by releasing and purifying the cellulose. Laccases are extremely important biocatalysts, which can be exploited for:
- paper pulp bleaching,
- detoxification (particularly bioremediation of pulp mill effluents),
- fibre modification,
- dye decolourisation,
- removal of phenolics from wines,
- organic synthesis and chemical transformation of pharmaceuticals (Sharma et al., 2018).
- as well as in self-regenerating microbial-enzymatic fuel cells (Evans et al., 2021).
Further, many pesticidal treatments depend on benzene rings for their effectiveness; examples are chemicals like pentachlorophenol (PCP) and polychlorinated biphenyls (PCBs). These persist in the environment because there are so few organisms able to catabolise them. But organisms that can catabolise lignin have all the tools necessary to destroy such persistent pesticides (Peralta et al. 2017).
DyP-type peroxidases and laccases are the major lignin-modifying enzymes produced by bacteria (de Gonzalo et al. 2016). Lignocellulose-consuming animals (termites, shipworms) secrete some glycoside hydrolases, but most harbour a diverse gut microbiota that includes fungi (chytrids) oomycetes and bacteria able to secrete lignocellulolytic enzymes in a mutualism that is particularly complex in termites (Cragg et al. 2015).
There are two intriguing side issues to fungal degradation of timber. One, which may become more crucial in the future, is that some of the small organic molecules the fungi produce to enhance activity of the lignin and manganese peroxidases incorporate chlorine from the wood into chloroaromatics, and even the synthesis of veratryl alcohol requires chloromethane. These volatile compounds are released into the atmosphere (thereby flushing the chlorine out of the substrate) and may themselves act as environmental pollutants. Some white-rot genera, particularly Phellinus and Inonotus, release enormous quantities of chloromethane into the atmosphere as they digest wood. The annual global release to the atmosphere from this source has been estimated at 160,000 tonnes, 75% of which is released from tropical and subtropical forests, with 86% being attributable to Phellinus spp. alone. Chloromethane is a powerful greenhouse gas and atmospheric pollutant which can have adverse effects on stratospheric ozone, yet in this case it is the product of a natural ecosystem (Watling & Harper, 1998; Anke & Weber, 2006).
The second intriguing aspect is that there are at least 75 species of fungi that are bioluminescent; they emit light from their mycelium and/or fruit bodies, and all species found so far have proved to be wood decay fungi. All known truly bioluminescent fungi are white rot fungi belonging to the order Agaricales (Basidiomycota) with one exceptional ascomycete belonging to the order Xylariales. So, they are mostly mushroom-forming, saprotrophic or, rarely, plant pathogenic species belonging to three distinct evolutionary lineages, which are named for their characteristic genera: 12 species belong to the Omphalotus lineage, five species to the Armillaria lineage (Armillaria mellea is the most widely distributed bioluminescent fungus), and 47 species belong to the mycenoid lineage. Different luminous fungi emit light from different tissues, for example, from:
- mycelium and rhizomorphs in Armillaria;
- mycelium only in many Mycena species;
- spores only in Mycena rorida;
- sclerotia only of Collybia tuberosa;
- mycelium and the whole (mushroom) fruit body in Panellus stipticus and Omphalotus olearius (= Clitocybe illudens); in the latter, light is emitted from the mushroom cap, stem, gills and spores.
Light emission in vitro can be obtained enzymatically by mixing cold extracts (which extract the enzymes) and hot extracts (which extract the substrates) from different species of fungi, which indicates a common mechanism for all these fungi. Kinetic data suggest a consecutive two-step enzymatic mechanism: first, a light emitting substrate (arbitrarily called ‘luciferin’) is reduced by a soluble reductase at the expense of NAD(P)H; second, reduced luciferin is oxidised by an insoluble (possibly membrane-bound?) luciferase that releases the energy in the form of bluish-green visible light with an emission maximum wavelength close to 530 nm.
Although fungal ‘luciferin’ is clearly one of the numerous secondary metabolites fungi can synthesise, its exact identity and structure has not yet been established but it is thought that all bioluminescent fungi share the same enzymatic mechanism, suggesting that the bioluminescent pathway arose early in the evolution of the Agaricales (Oliveira et al. 2012). Nor is it entirely clear what the physiological and ecological function of fungal bioluminescence might be (which makes it difficult to appreciate its evolutionary selective advantage). It has been suggested that in the dark beneath closed tropical forest canopies bioluminescent fruit bodies may be at an advantage by attracting grazing animals (and that would include insects and other arthropods) that could help disperse their spores. Conversely, where mycelium (and vegetative structures like rhizomorphs and sclerotia) are the bioluminescent tissues, the argument has been made that light emission could deter grazing. Neither of these suggestions is entirely satisfactory. However, as far as is known at present, all luminescent basidiomycetes are white-rot fungi capable of lignin degradation. Bioluminescence is an oxygen-dependent metabolic process and may be used for detoxifcation of peroxides formed during ligninolysis; with the energy being released as light rather than heat. The favoured hypothesis now, therefore, is that fungal bioluminescence is an advantageous process because it provides antioxidant protection against the potential deleterious effects of reactive oxygen species produced during wood decay (Desjardin et al., 2008; Audrey et al., 2015). Many mushrooms have been reported to possess antioxidant properties, which enable them to neutralise free radicals, and have important nutritional and therapeutic properties (Sánchez, 2017; Acosta-Urdapilleta et al., 2020).
Updated May, 2021
