9.2 Mycelial differentiation
Rhythmic, or cyclical, growth of colonies on solid medium is an excellent example of this interaction between the environment and the genetic capability of the mycelium. Regular concentric banding of colonies grown in vitro is seen quite often in many different fungi. It results from regular changes in hyphal extension rate and branch formation as hyphae react to local conditions as part of an endogenous or externally-regulated circadian clock (a roughly 24-hour cycle). These circadian rhythms are generated endogenously; under constant environmental conditions they are self-sustaining, and their period is determined genetically. However, they can be modulated by changes in external cues such as light, temperature and nutrients. Temporal rhythms are ubiquitous in eukaryotes and common regulatory patterns in circadian systems extend from fungi through to mammals. The formal study of biological rhythms (that may be daily, weekly, seasonal, annual or with an even longer period) is called chronobiology and covers an enormous range of animal (including human) and plant physiology. But cyclical changes are clearly evident in fungi and their causes are frequently more accessible than they are in other eukaryotes. We present a brief overview of the molecular aspects of circadian rhythms in the next few paragraphs.
Circadian rhythms are biological rhythms with periods of about 24 hours. Circadian clocks are molecular circuits that allow organisms to coordinate many processes, including gene expression, with a rhythm that is close to the daily 24-hour cycle. Rhythmic processes described in fungi include growth rate, stress responses, developmental capacity, and sporulation, as well as many metabolic processes. Generally, fungi use clocks to anticipate daily environmental changes. Rhythmicity is endogenous and self-sustaining when environmental conditions are constant; the length of the rhythmic cycle being genetically determined. Rhythmically changing environmental signals, particularly of light and temperature, set the phase of the endogenous rhythm and adjust it to exactly 24 hours.
Circadian clocks are self-sustaining timekeepers found in almost all organisms on earth. They have arisen at least three times through evolution, in prokaryotic cyanobacteria, in cells that evolved into higher plants, and in the opisthokont clade, the group of organisms that eventually became the fungi and the animals (Dunlap & Loros, 2017). They do not require complex tissue organisation, and even single cells can express rhythmicity. A negative feedback loop comprises the core of the circadian system in fungi and animals, centred on the transcription of clock genes and translation of clock proteins.
The positive element in the loop is the transcriptional activation of one or more clock gene(s) through binding of paired transcriptional activators on the clock gene promoter; they are paired by interaction through PAS domains (see below). Translation of the transcribed message of the clock gene (which is subject to additional regulation) generates a clock protein that is the negative element of the feedback loop. This blocks activation of the clock gene so the amount of clock gene mRNA declines and eventually the levels of clock protein also decline. These processes generate a daily cycle of clock gene mRNA and clock proteins and forms an oscillator that creates what is known as an ‘output’ that is the basis of the timing signal that controls rhythmical cellular functions (which might be organism-, organ- or even cell-specific) (Fig. 1). A circadian system can be made up of one or more interconnected feedback loops forming quite a complex network. In addition, the system receives inputs of ambient light and temperature to adjust its phase so that the internal day matches the external day, and then uses the time information it generates to regulate the life of the cell and of the whole organism.
 |
|---|
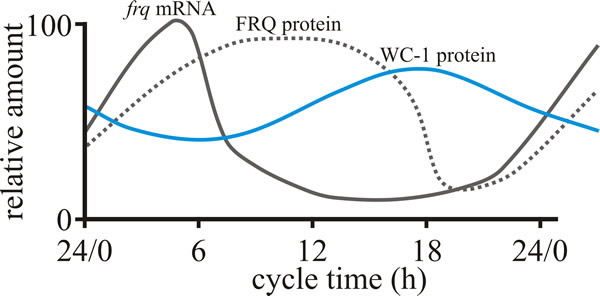
| Fig. 1. Behaviour of the FRQ –WC-1 based oscillator of Neurospora crassa over the course of a circadian cycle. The graph shows the daily cycles in levels of, FRQ protein and WC-1 protein (product of the white collar-1 gene) in constant darkness. Light activates production of frq mRNA, but ultimately the presence of excess FRQ protein represses this. Modified from Dunlap & Loros, 2006; and see Dunlap & Loros, 2017). |
Many clock gene proteins have a common structural motif known as the PAS domain. The name PAS is an acronym created from ‘PER-ARNT-Sim’ and PAS domains were first identified in the Drosophila proteins PER and ARNT and they were later found in a wide range of organisms. PAS domains are involved in many signalling proteins where they are used as a signal sensor domain. In circadian rhythmicity the PAS-domain proteins act as heterodimeric transcriptional activation complexes to drive expression of clock genes. Interestingly, though, PAS domains mediate protein-protein interactions in response to stimuli when cofactors bind within their hydrophobic cores, so they have important roles as sensory modules for a wide range of environmental conditions including oxygen tension, redox potential, carbon monoxide and nitric oxide, as well as light intensity and temperature; all of which are the main inputs to the Neurospora circadian oscillators.
The circadian clock of Neurospora, still the best studied fungal model, involves proteins (which are transcription factors) of mutant genes called white collar-1 (the protein is called WC-1), white collar-2 (WC-2), and frequency (gene symbol frq) (Koritala & Lee, 2017). FRQ (the protein encoded by the frq gene) is the core clock component (Cha et al., 2015) and complexes with other proteins, physically interacting with the WC transcription factors reducing their activity; the kinetics being strongly influenced by progressive phosphorylation of FRQ. When FRQ becomes sufficiently phosphorylated that it loses the ability to influence WC activities, the circadian cycle starts again. Environmental cycles of light and temperature influence frq and FRQ expression and thereby reset the internal circadian clocks (Dunlap & Loros, 2017). Light acts in Neurospora to induce transcription of the negative elements that reset the clock and synchronise the cell to the daily light/dark cycle. Temperature-influenced translational regulation of FRQ synthesis in Neurospora sets the physiological temperature limits over which the clock operates.
The circadian system of Neurospora is an important model system (Montenegro-Montero et al., 2015) used to understand circadian rhythms in other organisms. There is evidence for conservation of rhythmicity mechanisms in filamentous fungi in general; when tested for homology with Neurospora FRQ, WC-1 and WC-2 sequences, scores for similarity were high in genomes of Basidiomycota, and zygomycetes as well as other Ascomycota (Dunlap & Loros, 2006; 2017).
The Neurospora circadian system contains at least three oscillators:
- the FRQ/WC-dependent circadian oscillator, the core components of which are FRQ, WC-1, WC-2, and two other proteins, FRH and FWD-1;
- the WC-dependent circadian oscillator;
- and one or more FRQ/WC-independent oscillators.
A survey of 64 fungal proteomes for homologues of Neurospora clock proteins found that the FRH and FWD-1 proteins were probably present in the last common ancestor of all the fungi surveyed. Homologues of WC-1 and WC-2 were absent from chytrids and Microsporidia but were present in all other major clades. In contrast, FRQ homologues were restricted taxonomically within the Ascomycota to Sordariomycetes, Leotiomycetes and Dothideomycetes (Salichos & Rokas, 2010).
Our interpretation of these findings is that the components of the Neurospora circadian clock are widely conserved in fungal evolution, but the way they are assembled into a working oscillator in Neurospora is only one of several possibilities. The regulators that make up the clocks are involved in other cellular control events; for example, one of the FRQ homologues in the fungus Botrytis cinerea regulates virulence when the fungus is infecting its plant host Arabidopsis thaliana (Hevia et al., 2015; 2016). So, the development and evolution of a clock circuit will depend on the balance between the different selection pressures exerted on the different functions of its component parts.
We present a brief simplified overview of the molecular aspects of circadian rhythms in the Rhythms, oscillators and clock genes Resources Box, and from here concentrate on the mycelium morphology aspects.
Resources Box Rhythms, oscillators and clock genes Circadian rhythms are biological rhythms with periods of about 24 hours. Rhythmicity is endogenous and self-sustaining when environmental conditions are constant; the length of the rhythmic cycle being genetically determined. CLICK HERE to visit a page providing details of these. |
How rhythmic growth can influence mycelial morphology is illustrated by a clock-mutant of Podospora anserina which forms concentric bands of aerial growth within the colony grown on agar media. The banding arises from a difference in growth pattern of aerial and submerged hyphae. Enhanced growth and increased branching of hyphae on the agar surface eventually cause growth of aerial hyphae to stop, perhaps because of the accumulation of excretory products (known as ‘staling substances’), so further extension on the surface is limited by this ‘induction event’. On the other hand, submerged hyphae do not show this pattern of increased branching; they escape the restriction to growth and continue to extend and reach the surface some distance beyond the stopped surface mycelial front (Fig. 2).
Emergence of the submerged hyphae prevents further growth of the old surface mycelium but produces a new generation of surface hyphal tips which go through the same process of branching, staling and growth limitation. Repetition of the cycle gives rise to zones of alternately dense and sparse surface mycelium which are visible as a regular series of bands on the surface. Reduced extension rates and increased branching in this mycelium of Podospora anserina are accompanied by increased oxygen uptake and exposure to light.
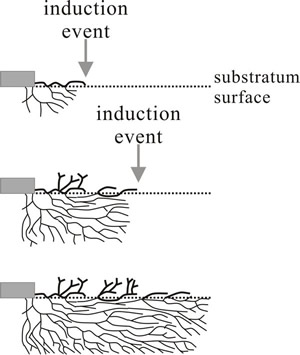 |
Fig. 2. Schematic illustration of the origin of rhythmic growth in a mutant of Podospora. Top: aerial and submerged hyphae grow away from the inoculum (grey rectangle) but in response to an induction event (may be an endogenous physiological cycle or external signal, like light or temperature fluctuation) surface growth is stopped. The check to surface extension prompts aerial branching and allows submerged hyphae to grow beyond the surface mycelium, branches from submerged hyphae re-establishing surface growth. A second induction event initiates another cycle of aerial branching after the check to extension on the surface (bottom diagram). Redrawn after Lysek, 1984. |
This example makes it clear that rhythmic growth is a differentiation process which separates hyphae with different functions and properties in space and time. In this case, extending hyphae, exploring for new substrates, are separated spatially from stationary surface hyphae, which may differentiate into sporing and/or resting structures after their extension growth is stopped.
Concentric rings and radial zonations of the mycelium are common expressions of mycelial growth rhythms. Evidently, an induction event must be detected and this suggests that membrane sensors play an important role in these sorts of reactions. The evidence indicates that many different sensory modules reacting to a wide range of environmental conditions (gases, redox, light, temperature) can input signals to a common oscillator that generates the rhythmic output (see the Rhythms, oscillators and clock genes Resources Box). The important point is that the rhythmic output is sent to the filamentous cell cycle discussed in Chapters 4 and 5, so that the essentially homogeneous growth and branching pattern of the vegetative hyphae is disturbed by the inducing event and differentiation of the mycelium results. This may be an intrahyphal differentiation, as in spore differentiation, or a concerted co-differentiation of several or many hyphae to produce the equivalent of a differentiated tissue, as in the concentric rings and zonations we have just been discussing.
Even in more complex fungal structures, like fruiting bodies, similar mechanisms may exist to change the hyphal growth and branching patterns to form functionally-distinct fungal tissues. Certainly, experiments based on transferring such tissues to artificial environments in vitro indicate that hyphal cells differentiate in response to signals they receive from their immediate environment within the ‘home’ tissue and that maintenance of their state of differentiation requires continual reinforcement from those signals (Chiu & Moore, 1988a & b). It is what those signals do that is important, and the general concept is that signals that control fungal differentiation do so by modifying the pattern (that is, the temporal and/or spatial distribution) of any or all of the numerous components that contribute to the fungal cell cycle. Methods are being developed to apply next-generation physiology approaches to study microbial communities at the single cell level (Hatzenpichler et al., 2020). If these could be applied to a filamentous fungal hypha, they could catalogue how successive compartments behave in normal extension growth and during differentiation, and how different hyphal septa function in this.
Resources Box Hyphal differentiation Differentiation of hyphae gives rise to an almost infinite variety of hyphal and cell shapes that have:
The prime source for explanation of names and terms in mycology is the Dictionary of the Fungi (Kirk et al., 2001), and a good alternative is the Illustrated Dictionary of Mycology (Ulloa & Hanlin, 2000). In this Resources Box we use extracts from the Dictionary of the Fungi to show you both something of the range of diversity that has been observed in fungal cells and the usefulness of this reference source. CLICK HERE to visit a page providing details of these. |
Examine the examples shown in the Hyphal Differentiation Resources Box carefully to form an understanding of the way the shape of the fungal cell can be varied by varying the formation of wall. The main purpose of the Hyphal Differentiation Resources Box is to introduce you to the diversity of spore and hyphal morphology, and to the diverse terminology developed by mycologists to describe this. However, you should take the opportunity to examine the range of morphologies shown, so that you can understand how shape of a spore or hypha can be determined by precise control over the timing and position of renewed wall growth varying the formation of wall and associated machinery.
Updated September, 2020
